Molecular characterization and functional analysis of the Schistosoma mekongi Ca2+-dependent cysteine protease (calpain)
- PMID: 31362766
- PMCID: PMC6668146
- DOI: 10.1186/s13071-019-3639-9
Molecular characterization and functional analysis of the Schistosoma mekongi Ca2+-dependent cysteine protease (calpain)
Abstract
Background: Schistosoma mekongi, which causes schistosomiasis in humans, is an important public health issue in Southeast Asia. Treatment with praziquantel is the primary method of control but emergence of praziquantel resistance requires the development of alternative drugs and vaccines. Calcium-dependent cysteine protease (calpain) is a novel vaccine candidate that has been studied in S. mansoni, S. japonicum, and protozoans including malaria, leishmania and trypanosomes. However, limited information is available on the properties and functions of calpain in other Schistosoma spp., including S. mekongi. In this study, we functionally characterized calpain 1 of S. mekongi (SmeCalp1).
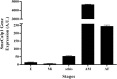
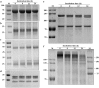
Results: Calpain 1 of S. mekongi was obtained from transcriptomic analysis of S. mekongi; it had the highest expression level of all isoforms tested and was predominantly expressed in the adult male. SmeCalp1 cDNA is 2274 bp long and encodes 758 amino acids, with 85% to 90% homology with calpains in other Schistosoma species. Recombinant SmeCalp1 (rSmeCalp1), with a molecular weight of approximately 86.7 kDa, was expressed in bacteria and stimulated a marked antibody response in mice. Native SmeCalp1 was detected in crude worm extract and excretory-secretory product, and it was mainly localized in the tegument of the adult male; less signal was detected in the adult female worm. Thus, SmeCalp1 may play a role in surface membrane synthesis or host-parasite interaction. We assessed the protease activity of rSmeCalp1 and demonstrated that rSmeCalp1 could cleave the calpain substrate N-succinyl-Leu-Leu-Val-Tyr-7-amino-4-methylcoumarin, that was inhibited by calpain inhibitors (MDL28170 and E64c). Additionally, rSmeCalp1 could degrade the biological substrates fibronectin (blood clotting protein) and human complement C3, indicating important roles in the intravascular system and in host immune evasion.
Conclusions: SmeCalp1 is expressed on the tegumental surface of the parasite and can cleave host defense molecules; thus, it might participate in growth, development and survival during the entire life-cycle of S. mekongi. Information on the properties and functions of SmeCalp1 reported herein will be advantageous in the development of effective drugs and vaccines against S. mekongi and other schistosomes.
Keywords: Calcium-dependent cysteine protease; Calpain; Drug and vaccine development; Schistosoma mekongi; Schistosomiasis.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures






Similar articles
-
Proteomic analysis of adult Schistosoma mekongi somatic and excretory-secretory proteins.Acta Trop. 2020 Feb;202:105247. doi: 10.1016/j.actatropica.2019.105247. Epub 2019 Oct 28. Acta Trop. 2020. PMID: 31672487
-
Proteomic and immunomic analysis of Schistosoma mekongi egg proteins.Exp Parasitol. 2018 Aug;191:88-96. doi: 10.1016/j.exppara.2018.07.002. Epub 2018 Jul 17. Exp Parasitol. 2018. PMID: 30009810
-
The blood fluke Schistosoma mansoni cleaves the coagulation protein high molecular weight kininogen (HK) but does not generate the vasodilator bradykinin.Parasit Vectors. 2018 Mar 14;11(1):182. doi: 10.1186/s13071-018-2704-0. Parasit Vectors. 2018. PMID: 29540224 Free PMC article.
-
Arachidonic Acid Is a Safe and Efficacious Schistosomicide, and an Endoschistosomicide in Natural and Experimental Infections, and Cysteine Peptidase Vaccinated Hosts.Front Immunol. 2020 Nov 17;11:609994. doi: 10.3389/fimmu.2020.609994. eCollection 2020. Front Immunol. 2020. PMID: 33281832 Free PMC article. Review.
-
Schistosomiasis mekongi: from discovery to control.Parasitol Int. 2004 Jun;53(2):135-42. doi: 10.1016/j.parint.2004.01.004. Parasitol Int. 2004. PMID: 15081945 Review.
Cited by
-
Transmission-Blocking Vaccines against Schistosomiasis Japonica.Int J Mol Sci. 2024 Jan 30;25(3):1707. doi: 10.3390/ijms25031707. Int J Mol Sci. 2024. PMID: 38338980 Free PMC article. Review.
-
Structural Transitions of Papain-like Cysteine Proteases: Implications for Sensor Development.Biomimetics (Basel). 2023 Jul 1;8(3):281. doi: 10.3390/biomimetics8030281. Biomimetics (Basel). 2023. PMID: 37504169 Free PMC article.
-
Characteristics and immunoprotective functions of three cysteine proteases from Clonorchis sinensis.Front Immunol. 2025 Apr 3;16:1550775. doi: 10.3389/fimmu.2025.1550775. eCollection 2025. Front Immunol. 2025. PMID: 40248698 Free PMC article.
-
Eudiplozoon nipponicum (Monogenea, Diplozoidae) and its adaptation to haematophagy as revealed by transcriptome and secretome profiling.BMC Genomics. 2021 Apr 15;22(1):274. doi: 10.1186/s12864-021-07589-z. BMC Genomics. 2021. PMID: 33858339 Free PMC article.
-
Secreted and surface proteome and transcriptome of Opisthorchis felineus.Front Parasitol. 2023 Oct 10;2:1195457. doi: 10.3389/fpara.2023.1195457. eCollection 2023. Front Parasitol. 2023. PMID: 39816815 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

