Identifying loci under positive selection in complex population histories
- PMID: 31362936
- PMCID: PMC6724678
- DOI: 10.1101/gr.246777.118
Identifying loci under positive selection in complex population histories
Abstract
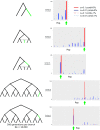
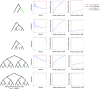
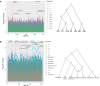
Detailed modeling of a species' history is of prime importance for understanding how natural selection operates over time. Most methods designed to detect positive selection along sequenced genomes, however, use simplified representations of past histories as null models of genetic drift. Here, we present the first method that can detect signatures of strong local adaptation across the genome using arbitrarily complex admixture graphs, which are typically used to describe the history of past divergence and admixture events among any number of populations. The method-called graph-aware retrieval of selective sweeps (GRoSS)-has good power to detect loci in the genome with strong evidence for past selective sweeps and can also identify which branch of the graph was most affected by the sweep. As evidence of its utility, we apply the method to bovine, codfish, and human population genomic data containing panels of multiple populations related in complex ways. We find new candidate genes for important adaptive functions, including immunity and metabolism in understudied human populations, as well as muscle mass, milk production, and tameness in specific bovine breeds. We are also able to pinpoint the emergence of large regions of differentiation owing to inversions in the history of Atlantic codfish.
© 2019 Refoyo-Martínez et al.; Published by Cold Spring Harbor Laboratory Press.
Figures




References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
