Human placenta has no microbiome but can contain potential pathogens
- PMID: 31367035
- PMCID: PMC6697540
- DOI: 10.1038/s41586-019-1451-5
Human placenta has no microbiome but can contain potential pathogens
Erratum in
-
Author Correction: Human placenta has no microbiome but can contain potential pathogens.Nature. 2019 Oct;574(7778):E15. doi: 10.1038/s41586-019-1628-y. Nature. 2019. PMID: 31570886
Abstract
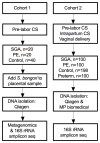
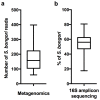
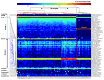
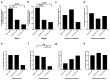
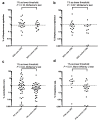
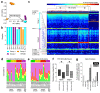
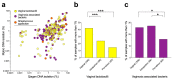
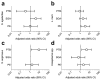
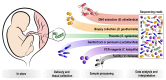
We sought to determine whether pre-eclampsia, spontaneous preterm birth or the delivery of infants who are small for gestational age were associated with the presence of bacterial DNA in the human placenta. Here we show that there was no evidence for the presence of bacteria in the large majority of placental samples, from both complicated and uncomplicated pregnancies. Almost all signals were related either to the acquisition of bacteria during labour and delivery, or to contamination of laboratory reagents with bacterial DNA. The exception was Streptococcus agalactiae (group B Streptococcus), for which non-contaminant signals were detected in approximately 5% of samples collected before the onset of labour. We conclude that bacterial infection of the placenta is not a common cause of adverse pregnancy outcome and that the human placenta does not have a microbiome, but it does represent a potential site of perinatal acquisition of S. agalactiae, a major cause of neonatal sepsis.
Conflict of interest statement
JP reports grants from Pfizer, personal fees from Next Gen Diagnostics Llc, outside the submitted work; SJP reports personal fees from Specific, personal fees from Next Gen Diagnostics, outside the submitted work; DSC-J reports grants from GlaxoSmithKline Research and Development Limited, outside the submitted work and non-financial support from Roche Diagnostics Ltd, outside the submitted work; GCSS reports grants and personal fees from GlaxoSmithKline Research and Development Limited, personal fees and non-financial support from Roche Diagnostics Ltd, outside the submitted work; DSC-J and GCSS report grants from Sera Prognostics Inc, non-financial support from Illumina inc, outside the submitted work. MCG, SL, US, FG and EC have nothing to disclose.
Figures




Comment in
-
No bacteria found in healthy placentas.Nature. 2019 Aug;572(7769):317-318. doi: 10.1038/d41586-019-02262-8. Nature. 2019. PMID: 31406307 No abstract available.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

