An integrated chromosome-scale genome assembly of the Masai giraffe (Giraffa camelopardalis tippelskirchi)
- PMID: 31367745
- PMCID: PMC6669057
- DOI: 10.1093/gigascience/giz090
An integrated chromosome-scale genome assembly of the Masai giraffe (Giraffa camelopardalis tippelskirchi)
Abstract
Background: The Masai giraffe (Giraffa camelopardalis tippelskirchi) is the largest-bodied giraffe and the world's tallest terrestrial animal. With its extreme size and height, the giraffe's unique anatomical and physiological adaptations have long been of interest to diverse research fields. Giraffes are also critical to ecosystems of sub-Saharan Africa, with their long neck serving as a conduit to food sources not shared by other herbivores. Although the genome of a Masai giraffe has been sequenced, the assembly was highly fragmented and suboptimal for genome analysis. Herein we report an improved giraffe genome assembly to facilitate evolutionary analysis of the giraffe and other ruminant genomes.
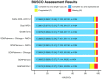
Findings: Using SOAPdenovo2 and 170 Gbp of Illumina paired-end and mate-pair reads, we generated a 2.6-Gbp male Masai giraffe genome assembly, with a scaffold N50 of 3 Mbp. The incorporation of 114.6 Gbp of Chicago library sequencing data resulted in a HiRise SOAPdenovo + Chicago assembly with an N50 of 48 Mbp and containing 95% of expected genes according to BUSCO analysis. Using the Reference-Assisted Chromosome Assembly tool, we were able to order and orient scaffolds into 42 predicted chromosome fragments (PCFs). Using fluorescence in situ hybridization, we placed 153 cattle bacterial artificial chromosomes onto giraffe metaphase spreads to assess and assign the PCFs on 14 giraffe autosomes and the X chromosome resulting in the final assembly with an N50 of 177.94 Mbp. In this assembly, 21,621 protein-coding genes were identified using both de novo and homology-based predictions.
Conclusions: We have produced the first chromosome-scale genome assembly for a Giraffidae species. This assembly provides a valuable resource for the study of artiodactyl evolution and for understanding the molecular basis of the unique adaptive traits of giraffes. In addition, the assembly will provide a powerful resource to assist conservation efforts of Masai giraffe, whose population size has declined by 52% in recent years.
Keywords: Giraffa camelopardalis tippelskirchi; annotation; assembly; giraffe; ruminant.
© The Author(s) 2019. Published by Oxford University Press.
Figures



References
-
- Fennessy J, Bidon T, Reuss F, et al. .. Multi-locus analyses reveal four giraffe species instead of one. Curr Biol. 2016;26(18):2543–9. - PubMed
-
- Muller Z, Bercovitch F, Brand R et al. .. Giraffa camelopardalis. The IUCN Red List of Threatened Species 2016, United Kingdom. 2016, 10.2305/IUCN.UK.2016-3.RLTS.T9194A136266699.en. - DOI
-
- Dagg AI. Giraffe: Biology, Behaviour and Conservation. Cambridge, UK: Cambridge University Press; 2014.
-
- Solounias N. The remarkable anatomy of the giraffe's neck. J Zool. 1999;247(2):257–68.
-
- Estes R. The Behavior Guide to African Mammals: Including Hoofed Mammals, Carnivores, Primates. Berkeley: University of California Press; 1991.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

