De novo assembly of a wild pear (Pyrus betuleafolia) genome
- PMID: 31368610
- PMCID: PMC6953202
- DOI: 10.1111/pbi.13226
De novo assembly of a wild pear (Pyrus betuleafolia) genome
Abstract
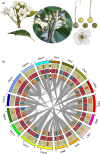
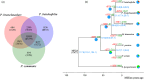
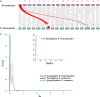
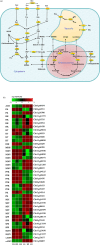
China is the origin and evolutionary centre of Oriental pears. Pyrus betuleafolia is a wild species native to China and distributed in the northern region, and it is widely used as rootstock. Here, we report the de novo assembly of the genome of P. betuleafolia-Shanxi Duli using an integrated strategy that combines PacBio sequencing, BioNano mapping and chromosome conformation capture (Hi-C) sequencing. The genome assembly size was 532.7 Mb, with a contig N50 of 1.57 Mb. A total of 59 552 protein-coding genes and 247.4 Mb of repetitive sequences were annotated for this genome. The expansion genes in P. betuleafolia were significantly enriched in secondary metabolism, which may account for the organism's considerable environmental adaptability. An alignment analysis of orthologous genes showed that fruit size, sugar metabolism and transport, and photosynthetic efficiency were positively selected in Oriental pear during domestication. A total of 573 nucleotide-binding site (NBS)-type resistance gene analogues (RGAs) were identified in the P. betuleafolia genome, 150 of which are TIR-NBS-LRR (TNL)-type genes, which represented the greatest number of TNL-type genes among the published Rosaceae genomes and explained the strong disease resistance of this wild species. The study of flavour metabolism-related genes showed that the anthocyanidin reductase (ANR) metabolic pathway affected the astringency of pear fruit and that sorbitol transporter (SOT) transmembrane transport may be the main factor affecting the accumulation of soluble organic matter. This high-quality P. betuleafolia genome provides a valuable resource for the utilization of wild pear in fundamental pear studies and breeding.
Keywords: Pyrus betuleafolia; BioNano optical mapping; De novo assembly; Hi-C; PacBio SMRT.
© 2019 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
No conflicts of interest are declared.
Figures


Similar articles
-
A de novo genome assembly of the dwarfing pear rootstock Zhongai 1.Sci Data. 2019 Nov 25;6(1):281. doi: 10.1038/s41597-019-0291-3. Sci Data. 2019. PMID: 31767847 Free PMC article.
-
The genome of the pear (Pyrus bretschneideri Rehd.).Genome Res. 2013 Feb;23(2):396-408. doi: 10.1101/gr.144311.112. Epub 2012 Nov 13. Genome Res. 2013. PMID: 23149293 Free PMC article.
-
Pseudo-chromosome-length genome assembly of a double haploid "Bartlett" pear (Pyrus communis L.).Gigascience. 2019 Dec 1;8(12):giz138. doi: 10.1093/gigascience/giz138. Gigascience. 2019. PMID: 31816089 Free PMC article.
-
The draft genome sequence of European pear (Pyrus communis L. 'Bartlett').PLoS One. 2014 Apr 3;9(4):e92644. doi: 10.1371/journal.pone.0092644. eCollection 2014. PLoS One. 2014. PMID: 24699266 Free PMC article.
-
Genetic structure and diversity of the wild Ussurian pear in East Asia.Breed Sci. 2016 Jan;66(1):90-9. doi: 10.1270/jsbbs.66.90. Epub 2016 Jan 1. Breed Sci. 2016. PMID: 27069394 Free PMC article. Review.
Cited by
-
PbrWRKY62-PbrADC1 module involves in superficial scald development of Pyrus bretschneideri Rehd.fruit via regulating putrescine biosynthesis.Mol Hortic. 2024 Feb 20;4(1):6. doi: 10.1186/s43897-024-00081-8. Mol Hortic. 2024. PMID: 38373989 Free PMC article.
-
De novo assembly and characterization of the first draft genome of quince (Cydonia oblonga Mill.).Sci Rep. 2021 Feb 15;11(1):3818. doi: 10.1038/s41598-021-83113-3. Sci Rep. 2021. PMID: 33589687 Free PMC article.
-
Comparative genomic analyses reveal different genetic basis of two types of fruit in Maloideae.Nat Commun. 2025 Aug 12;16(1):7463. doi: 10.1038/s41467-025-62850-3. Nat Commun. 2025. PMID: 40796913 Free PMC article.
-
Genome-Wide Identification of Genes Encoding for Rho-Related Proteins in 'Duli' Pear (Pyrus betulifolia Bunge) and Their Expression Analysis in Response to Abiotic Stress.Plants (Basel). 2022 Jun 19;11(12):1608. doi: 10.3390/plants11121608. Plants (Basel). 2022. PMID: 35736759 Free PMC article.
-
The genome of kenaf (Hibiscus cannabinus L.) provides insights into bast fibre and leaf shape biogenesis.Plant Biotechnol J. 2020 Aug;18(8):1796-1809. doi: 10.1111/pbi.13341. Epub 2020 Jan 30. Plant Biotechnol J. 2020. PMID: 31975524 Free PMC article.
References
-
- Bailey, L.H. (1917) Pyrus. Standard cyclopedia of horticulture, vol. V, pp. 2865–2878. New York: Macmillan.
-
- Bell, R. , Quamme, H. , Layne, R. and Skirvin, R. (1996). Pears. In Fruit breeding, vol 1: tree and tropical fruits ( Janick, J. and Moore, J.N. , eds), pp. 441–514. New York: Wiley.
-
- Bouvier, L. , Bourcy, M. , Boulay, M. , Tellier, M. , Guérif, P. , Denancé, C. , Durel, C.E. et al. (2012) The new pear scab resistance gene Rvp1 from the European pear cultivar ‘Navara’ maps in a genomic region syntenic to an apple scab resistance gene cluster on linkage group 2. Tree Genet. Genomes, 8, 53–60.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

