The RNA hairpin binder TRIM71 modulates alternative splicing by repressing MBNL1
- PMID: 31371437
- PMCID: PMC6719626
- DOI: 10.1101/gad.328492.119
The RNA hairpin binder TRIM71 modulates alternative splicing by repressing MBNL1
Abstract
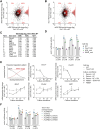
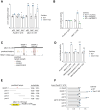
TRIM71/LIN-41, a phylogenetically conserved regulator of development, controls stem cell fates. Mammalian TRIM71 exhibits both RNA-binding and protein ubiquitylation activities, but the functional contribution of either activity and relevant primary targets remain poorly understood. Here, we demonstrate that TRIM71 shapes the transcriptome of mouse embryonic stem cells (mESCs) predominantly through its RNA-binding activity. We reveal that TRIM71 binds targets through 3' untranslated region (UTR) hairpin motifs and that it acts predominantly by target degradation. TRIM71 mutations implicated in etiogenesis of human congenital hydrocephalus impair target silencing. We identify a set of primary targets consistently regulated in various human and mouse cell lines, including MBNL1 (Muscleblind-like protein 1). MBNL1 promotes cell differentiation through regulation of alternative splicing, and we demonstrate that TRIM71 promotes embryonic splicing patterns through MBNL1 repression. Hence, repression of MBNL1-dependent alternative splicing may contribute to TRIM71's function in regulating stem cell fates.
Keywords: 3′ UTR; 5′ UTR; LIN41; RNA-binding protein; TRIM71; alternative splicing; mRNA degradation; muscleblind; stem cell; translational repression.
© 2019 Welte et al.; Published by Cold Spring Harbor Laboratory Press.
Figures





Similar articles
-
Molecular mechanism governing RNA-binding property of mammalian TRIM71 protein.Sci Bull (Beijing). 2024 Jan 15;69(1):72-81. doi: 10.1016/j.scib.2023.11.041. Epub 2023 Nov 21. Sci Bull (Beijing). 2024. PMID: 38036331
-
A congenital hydrocephalus-causing mutation in Trim71 induces stem cell defects via inhibiting Lsd1 mRNA translation.EMBO Rep. 2023 Feb 6;24(2):e55843. doi: 10.15252/embr.202255843. Epub 2022 Dec 27. EMBO Rep. 2023. PMID: 36573342 Free PMC article.
-
TRIM71 binds to IMP1 and is capable of positive and negative regulation of target RNAs.Cell Cycle. 2020 Sep;19(18):2314-2326. doi: 10.1080/15384101.2020.1804232. Epub 2020 Aug 20. Cell Cycle. 2020. PMID: 32816599 Free PMC article.
-
Molecular and biological functions of TRIM-NHL RNA-binding proteins.Wiley Interdiscip Rev RNA. 2021 Mar;12(2):e1620. doi: 10.1002/wrna.1620. Epub 2020 Aug 1. Wiley Interdiscip Rev RNA. 2021. PMID: 32738036 Free PMC article. Review.
-
The Muscleblind family of proteins: an emerging class of regulators of developmentally programmed alternative splicing.Differentiation. 2006 Mar;74(2-3):65-80. doi: 10.1111/j.1432-0436.2006.00060.x. Differentiation. 2006. PMID: 16533306 Review.
Cited by
-
TRIM71 Deficiency Causes Germ Cell Loss During Mouse Embryogenesis and Is Associated With Human Male Infertility.Front Cell Dev Biol. 2021 May 13;9:658966. doi: 10.3389/fcell.2021.658966. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34055789 Free PMC article.
-
The cytoplasmic fraction of the histone lysine methyltransferase Setdb1 is essential for embryonic stem cells.iScience. 2023 Jul 14;26(8):107386. doi: 10.1016/j.isci.2023.107386. eCollection 2023 Aug 18. iScience. 2023. PMID: 37559904 Free PMC article.
-
Mammalian RNA Decay Pathways Are Highly Specialized and Widely Linked to Translation.Mol Cell. 2020 Mar 19;77(6):1222-1236.e13. doi: 10.1016/j.molcel.2020.01.007. Epub 2020 Feb 10. Mol Cell. 2020. PMID: 32048998 Free PMC article.
-
Identification of RNA-binding proteins' direct effects on gene expression via the degradation tag system.RNA. 2023 Oct;29(10):1453-1457. doi: 10.1261/rna.079669.123. Epub 2023 Jul 6. RNA. 2023. PMID: 37414463 Free PMC article.
-
Impaired neurogenesis alters brain biomechanics in a neuroprogenitor-based genetic subtype of congenital hydrocephalus.Nat Neurosci. 2022 Apr;25(4):458-473. doi: 10.1038/s41593-022-01043-3. Epub 2022 Apr 4. Nat Neurosci. 2022. PMID: 35379995 Free PMC article.
References
-
- Batra R, Charizanis K, Manchanda M, Mohan A, Li M, Finn DJ, Goodwin M, Zhang C, Sobczak K, Thornton CA, et al. 2014. Loss of MBNL leads to disruption of developmentally regulated alternative polyadenylation in RNA-mediated disease. Mol Cell 56: 311–322. 10.1016/j.molcel.2014.08.027 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
