Identification of multiple potent neutralizing and non-neutralizing antibodies against Epstein-Barr virus gp350 protein with potential for clinical application and as reagents for mapping immunodominant epitopes
- PMID: 31377598
- PMCID: PMC6733660
- DOI: 10.1016/j.virol.2019.07.026
Identification of multiple potent neutralizing and non-neutralizing antibodies against Epstein-Barr virus gp350 protein with potential for clinical application and as reagents for mapping immunodominant epitopes
Abstract
Prevention of Epstein-Barr virus (EBV) infection has focused on generating neutralizing antibodies (nAbs) targeting the major envelope glycoprotein gp350/220 (gp350). In this study, we generated 23 hybridomas producing gp350-specific antibodies. We compared the candidate gp350-specific antibodies to the well-characterized nAb 72A1 by: (1) testing their ability to detect gp350 using enzyme-linked immunosorbent assay, flow cytometry, and immunoblot; (2) sequencing their heavy and light chain complementarity-determining regions (CDRs); (3) measuring the ability of each monoclonal antibody (mAb) to neutralize EBV infection in vitro; and (4) mapping the gp350 amino acids bound by the mAbs using competitive cell and linear peptide binding assays. We performed sequence analysis to identify 15 mAbs with CDR regions unique from those of murine 72A1 (m72A1). We observed antigen binding competition between biotinylated m72A1, serially diluted unlabeled gp350 nAbs (HB1, HB5, HB11, HB20), and our recently humanized 72A1, but not gp350 non-nAb (HB17) or anti-KSHV gH/gL antibody.
Keywords: Complementarity-determining region; Epitope; Epstein-Barr virus; Immunodominant; Immunosuppression; Infection; Neutralizing antibodies; gp350.
Copyright © 2019 Elsevier Inc. All rights reserved.
Conflict of interest statement
Figures
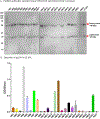





Similar articles
-
Peptides designed to spatially depict the Epstein-Barr virus major virion glycoprotein gp350 neutralization epitope elicit antibodies that block virus-neutralizing antibody 72A1 interaction with the native gp350 molecule.J Virol. 2015 May;89(9):4932-41. doi: 10.1128/JVI.03269-14. Epub 2015 Feb 18. J Virol. 2015. PMID: 25694592 Free PMC article.
-
High Epstein-Barr Virus Load and Genomic Diversity Are Associated with Generation of gp350-Specific Neutralizing Antibodies following Acute Infectious Mononucleosis.J Virol. 2016 Dec 16;91(1):e01562-16. doi: 10.1128/JVI.01562-16. Print 2017 Jan 1. J Virol. 2016. PMID: 27733645 Free PMC article.
-
Epstein-Barr Virus gp350 Can Functionally Replace the Rhesus Lymphocryptovirus Major Membrane Glycoprotein and Does Not Restrict Infection of Rhesus Macaques.J Virol. 2015 Nov 11;90(3):1222-30. doi: 10.1128/JVI.02531-15. Print 2016 Feb 1. J Virol. 2015. PMID: 26559839 Free PMC article.
-
Epstein Barr Virus: Development of Vaccines and Immune Cell Therapy for EBV-Associated Diseases.Front Immunol. 2021 Oct 8;12:734471. doi: 10.3389/fimmu.2021.734471. eCollection 2021. Front Immunol. 2021. PMID: 34691042 Free PMC article. Review.
-
Multiplicity of uses of monoclonal antibodies that define papillomavirus linear immunodominant epitopes.Immunol Res. 1997 Feb;16(1):115-9. doi: 10.1007/BF02786327. Immunol Res. 1997. PMID: 9048212 Review.
Cited by
-
Four Decades of Prophylactic EBV Vaccine Research: A Systematic Review and Historical Perspective.Front Immunol. 2022 Apr 14;13:867918. doi: 10.3389/fimmu.2022.867918. eCollection 2022. Front Immunol. 2022. PMID: 35493498 Free PMC article.
-
Multivalent MVA-vectored vaccine elicits EBV neutralizing antibodies in rhesus macaques that reduce EBV infection in humanized mice.Front Immunol. 2024 Sep 13;15:1445209. doi: 10.3389/fimmu.2024.1445209. eCollection 2024. Front Immunol. 2024. PMID: 39346922 Free PMC article.
-
Immunization with a self-assembling nanoparticle vaccine displaying EBV gH/gL protects humanized mice against lethal viral challenge.Cell Rep Med. 2022 Jun 21;3(6):100658. doi: 10.1016/j.xcrm.2022.100658. Epub 2022 Jun 14. Cell Rep Med. 2022. PMID: 35705092 Free PMC article.
-
Whole genome sequence analysis of equid gammaherpesvirus -2 field isolates reveals high levels of genomic diversity and recombination.BMC Genomics. 2022 Aug 30;23(1):622. doi: 10.1186/s12864-022-08789-x. BMC Genomics. 2022. PMID: 36042397 Free PMC article.
-
Evolution of functional antibodies following acute Epstein-Barr virus infection.PLoS Pathog. 2022 Sep 6;18(9):e1010738. doi: 10.1371/journal.ppat.1010738. eCollection 2022 Sep. PLoS Pathog. 2022. PMID: 36067220 Free PMC article.
References
-
- Alexander BT, Hladnik LM, Augustin KM, Casabar E, McKinnon PS, Reichley RM, Ritchie DJ, Westervelt P, Dubberke ER, 2010. Use of Cytomegalovirus Intravenous Immune Globulin for the Adjunctive Treatment of Cytomegalovirus in Hematopoietic Stem Cell Transplant Patients. Pharmacotherapy 30, 554–561. - PMC - PubMed
-
- Alfarano C, Andrade CE, Anthony K, Bahroos N, Bajec M, Bantoft K, Betel D, Bobechko B, Boutilier K, Burgess E, Buzadzija K, Cavero R, D’Abreo C, Donaldson I, Dorairajoo D, Dumontier MJ, Dumontier MR, Earles V, Farrall R, Feldman H, Garderman E, Gong Y, Gonzaga R, Grytsan V, Gryz E, Gu V, Haldorsen E, Halupa A, Haw R, Hrvojic A, Hurrell L, Isserlin R, Jack F, Juma F, Khan A, Kon T, Konopinsky S, Le V, Lee E, Ling S, Magidin M, Moniakis J, Montojo J, Moore S, Muskat B, Ng I, Paraiso JP, Parker B, Pintilie G, Pirone R, Salama JJ, Sgro S, Shan T, Shu Y, Siew J, Skinner D, Snyder K, Stasiuk R, Strumpf D, Tuekam B, Tao S, Wang Z, White M, Willis R, Wolting C, Wong S, Wrong A, Xin C, Yao R, Yates B, Zhang S, Zheng K, Pawson T, Ouellette BF, Hogue CW, 2005. The Biomolecular Interaction Network Database and related tools 2005 update. Nucleic acids research 33, D418–424. - PMC - PubMed
-
- Babcock GJ, Decker LL, Volk M, Thorley-Lawson DA, 1998. EBV persistence in memory B cells in vivo. Immunity 9, 395–404. - PubMed
-
- Baer R, Bankier AT, Biggin MD, Deininger PL, Farrell PJ, Gibson TJ, Hatfull G, Hudson GS, Satchwell SC, Seguin C, Tuffnell PS, Barrell BG, 1984. DNA-Sequence and Expression of the B95–8 Epstein-Barr Virus Genome. Nature 310, 207–211. - PubMed
-
- Benkerrou M, Jais JP, Leblond V, Durandy A, Sutton L, Bordigoni P, Garnier JL, Le Bidois J, Le Deist F, Blanche S, Fischer A, 1998. Anti-B-cell monoclonal antibody treatment of severe posttransplant B-lymphoproliferative disorder: prognostic factors and long-term outcome. Blood 92, 3137–3147. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

