Partially inserted nascent chain unzips the lateral gate of the Sec translocon
- PMID: 31379073
- PMCID: PMC6776908
- DOI: 10.15252/embr.201948191
Partially inserted nascent chain unzips the lateral gate of the Sec translocon
Abstract
The Sec translocon provides the lipid bilayer entry for ribosome-bound nascent chains and thus facilitates membrane protein biogenesis. Despite the appreciated role of the native environment in the translocon:ribosome assembly, structural information on the complex in the lipid membrane is scarce. Here, we present a cryo-electron microscopy-based structure of bacterial translocon SecYEG in lipid nanodiscs and elucidate an early intermediate state upon insertion of the FtsQ anchor domain. Insertion of the short nascent chain causes initial displacements within the lateral gate of the translocon, where α-helices 2b, 7, and 8 tilt within the membrane core to "unzip" the gate at the cytoplasmic side. Molecular dynamics simulations demonstrate that the conformational change is reversed in the absence of the ribosome, and suggest that the accessory α-helices of SecE subunit modulate the lateral gate conformation. Site-specific cross-linking validates that the FtsQ nascent chain passes the lateral gate upon insertion. The structure and the biochemical data suggest that the partially inserted nascent chain remains highly flexible until it acquires the transmembrane topology.
Keywords: membrane protein insertion; nanodisc; native environment; reconstitution; ribosome.
© 2019 The Authors. Published under the terms of the CC BY 4.0 license.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures

Structure of quiescent SecYEG of Thermus thermophilus in the lipid cubic phase (PDB ID: 5AWW). TMHs 2b, 3, 7, and 8 of the lateral gate, as well as the proximate loop 6/7 involved in ribosome binding are indicated. The non‐essential SecG subunit is omitted for clarity.
Model of the SecY lateral gate opening upon inserting a nascent chain (red) in the lipid bilayer. The color‐coding of SecYE TMHs is as in panel (A). In the presence of the completely inserted and folded nascent chain, TMHs 2b and 3 of the N‐terminal domain of SecY are displaced (arrows) thus opening a broad passage for the nascent TMH toward the lipid moiety.
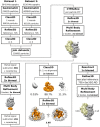
SDS–PAGE of SecYEG‐ND sample after size‐exclusion chromatography. Asterisks indicate translocon‐enriched fractions used for forming the RNC FtsQ:SecYEG‐ND complex. Lipid‐loaded “empty” nanodiscs elute at larger volumes and so can be separated.
Schematic drawing of a SecYEG‐ND particle. Lateral dimensions of the nanodisc should be appropriate to accommodate a single SecYEG with surroundings lipids, thus mimicking the naturally occurring environment.

The architecture of FtsQ nascent chain: N‐terminal hexa‐histidine tag was fused via 3C protease cleavage site to the N‐terminal domain of FtsQ followed by hemagglutinin (HA) recognition tag and TnaC stalling sequence. The length (amino acids) of each segment is indicated.
Coomassie‐stained protein content of isolated RNC FtsQ (left) and Western blot against the HA tag within the nascent chain (right). Western blot visualizes a complex of the FtsQ nascent chain (109 aa, ˜12 kDa) and tRNA (˜25 kDa).
Putative architecture of RNC:SecYEG‐ND complex upon the insertion of the early intermediate of FtsQ.
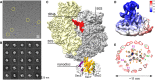
Representative cryo‐EM micrograph of RNC FtsQ:SecYEG‐ND. Exemplary individual ribosomes are encircled.
Examples of two‐dimensional classes of imaged particles. RNC:nanodisc assemblies can be seen at different view angles.
Three‐dimensional reconstruction of RNC FtsQ:SecYEG‐ND complex. Primary structural elements of the ribosome and SecYEG‐ND are indicated.
Local resolution map of SecYEG‐ND sub‐particle. The cytoplasmic side of the translocon demonstrates higher resolution due to stabilization by the bound ribosome, while high resolution at the periplasmic side is hindered by the SecYEG‐ND dynamics within the complex. The associated ribosome is not shown for clarity.
A planar slice through the SecYEG‐ND core at different signal levels (blue/green/red) with indicated positions of SecYEG TMHs (SecY indicated in orange, SecE in purple, and SecG in green). A single helical turn could be fitted in a density in the area where SecG TMH 1 was expected (green asterisk).

Isolated cryo‐EM density of SecYEG with the fitted molecular model of the translocon in front and the cytoplasmic views. SecY is displayed in rainbow pattern, SecE in purple, and SecG in green. Dashed box: a contact site between tilted SecE TMHs 1/2 and SecY TMH 8.
Cryo‐EM density corresponding to the ribosomal RNA helix 59 (“H59”) is displaced toward the nanodisc. No contact with the lipid bilayer could be detected. Blue ribbon: structure of the translocon‐free 50S ribosomal subunit (PDB ID: 4UY8).
Central cross‐section through the SecYEG model. The “plug” TMH 2a occupies the central position, thus keeping the translocon sealed. The nanodisc perimeter is indicated as a dashed circle.
The lateral gate of nanodisc‐embedded translocon undergoes rearrangements relatively to a quiescent conformation (left, PDB ID 5AWW) and an RNC‐bound detergent‐solubilized state (right, PDB ID 5GAE).
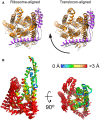
Superimposition of SecYEG‐ND (colored) and detergent‐solubilized translocon (gray, PDB ID: 5GAE) reveals different alignments if either 50S ribosome subunit (left) or the translocon itself (right) was used as an alignment template. Depending on the environment, the translocon undergoes a rotation around the ribosome‐bound loop 6/7.
The movement directions are indicated by rods, and the magnitude, as measured for positions of Cα atoms, by color (central panel).

- A–F
(A, D): Average all‐atom atomic fluctuations (RMSF; root‐mean‐square fluctuations) for SecY (A) and SecE (D). The mean RMSFs are projected onto the 3D structures of SecY/SecE and colored according to the color scales starting from blue (low mobility) to red (highly mobile). (B, E): Overlay of the SecY (B) or SecE (E) representative structure (red) onto the corresponding starting structure (dark cyan). Parts that show a pronounced structural change are explicitly shown as cartoon representation, and the movements are highlighted by arrows. (C, F): Visualization of displacements along the 1st (golden arrows) principal component computed for the joint, five 1 μs long MD simulations for SecY (C) and SecE (F). The amplitudes of the motions were scaled, and a cutoff for small displacements was applied for best graphical representation.
- G
Distribution of lipids during MD simulations. Grids represent the 3D density of phosphatidylcholine (PC; blue) and phosphatidylglycerol (PG; red) groups from the lipid bilayer. The densities were normalized to the number of considered conformations, which are identical in both cases.

An overview of intramolecular distances (I‐IV) measured to characterize the structural dynamics of the lateral gate of SecY. All distances were measured between the centers of mass (yellow spheres) of four residues that compose a helical turn (relevant residues are colored yellow and explicitly labeled; center of mass distances are indicated by the cyan, dashed lines).
Distance measurements from MD simulations. The left panels show the distances as a function of the simulation time for five MD simulations (differently colored lines) and right panels the mean relative frequencies (n = 5) plotted as a histogram. Gray areas denote the standard error of the mean for each bin. Cyan lines denote the distance in the starting structure.
Definition of the angle η that describes the relative orientation of SecY TMH 7 (“α7”) to TMH 8 (“α8”). The backbone centers of mass (yellow spheres) of the terminal four residues of TMHs 7 and 8 (colored yellow) served as the start and end points for the vectors (red arrows), respectively.
Relative motions of TMHs 7 and 8 in MD simulations. The left panel shows the angle η as a function of the simulation time and the right panel the mean relative frequencies (n = 5) plotted as a histogram. The gray area denotes to the standard error of the mean for each bin. The cyan line denotes the value of η in the starting structure.
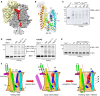
- A, B
Cryo‐EM visualizes tRNA‐bound nascent chain (red, NC) within the ribosomal tunnel (A) and a density between separated TMHs 2b and 7 of the lateral gate (B, shown in mesh). The proximate cysteine positions within SecY used for cross‐linking are indicated and highlighted in red.
- C
Western blot against the hemagglutinin tag within the nascent chain FtsQC40 reveals a cross‐linking product of ˜80 kDa in presence of nanodisc‐reconstituted SecYC282EG and SecYC283EG, presumably assigned to FtsQxSecY adduct. The double band for FtsQC40 likely indicates the incomplete cleavage of the hexa‐histidine tag by 3C protease.
- D
In‐gel fluorescence imaging of SecYCF488A reveals a cross‐linking product of ˜80 kDa assigned to the FtsQC40xSecYC283 adduct (“SecYxFtsQ”). Bands for covalently cross‐linked SecY dimer (“SecYxSecY”), and adducts of SecY and ribosomal proteins (“SecYxRiboP”) are indicated. The nascent chain‐specific adduct “SecYxFtsQ” does not depend on the presence of PE lipids.
- E
FtsQ:SecY cross‐linking is position specific. FtsQ TMH C‐terminal residues 40‐42 could be cross‐linked to the lateral gate (SecY residue 283), while no adduct could be detected at the upstream position 35.
- F
A refined scheme of the nascent chain insertion via the translocon. In the early intermediate state, the ribosome binding to the loop 6/7 and the emerging nascent chain cause displacements of SecY TMHs 2b and 7 at the cytoplasmic interface and “unzip” the lateral gate. The rigid‐body tilt of TMH 7 leads to a close contact between TMHs 3 and 7 at the periplasmic side, so the V‐shaped crevice is formed. The flexible nascent chain reaches the core of the lateral gate and gradually egresses the lipid moiety, where the helical fold is stabilized by hydrophobic interactions. Upon further nascent chain elongation and movements of SecY TMHs 2b, 3, 7, and 8, the widely open “inserting” state of the translocon is formed, with the complete nascent TMH at the lateral gate prior the release into the membrane.

A globular electron density centered above the SecY pore (circle) and a rod‐shaped density within the hollow nanodisc lumen opposite to the lateral gate (square) may represent the flexible FtsQ nascent chain.
Newly introduced cysteines within the lateral gate used for cross‐linking are deeply buried in the membrane and well‐distanced from the residue 148 at the periplasmic side of the translocon (circle) where a fluorescent dye can be conjugated.
CF488A‐maleimide cannot access the residue 282 within the hydrophobic core of the translocon, while an efficient labeling is achieved with BODIPY, Cy3, and AlexaFluor 633 maleimide conjugates. Structures of tested dyes are shown (right).
CF488A‐maleimide may be used for specific labeling of periplasm‐facing cysteine at position 148, while all tested positions within the lateral gate are not accessible.
Treatment of the cross‐linked samples with N‐ethylmaleimide (NEM) has no effect on SecYxFtsQ and SecYxSecY (asterisk) cross‐linking, but it strongly reduces the amount of adducts between SecY and ribosomal proteins (SecYxRiboP), suggesting that the latter contacts are unspecifically formed upon the denaturing in the SDS–PAGE buffer.
Comment in
-
A snapshot of membrane protein insertion.EMBO Rep. 2019 Oct 4;20(10):e49034. doi: 10.15252/embr.201949034. Epub 2019 Sep 13. EMBO Rep. 2019. PMID: 31518050 Free PMC article.
References
-
- Hessa T, Meindl‐Beinker NM, Bernsel A, Kim H, Sato Y, Lerch‐Bader M, Nilsson I, White SH, Von Heijne G (2007) Molecular code for transmembrane‐helix recognition by the Sec61 translocon. Nature 450: 1026–1030 - PubMed
-
- Van Den Berg B, Clemons WM, Collinson I, Modis Y, Hartmann E, Harrison SC, Rapoport TA (2004) X‐ray structure of a protein‐conducting channel. Nature 427: 36–44 - PubMed
-
- Tanaka Y, Sugano Y, Takemoto M, Mori T, Furukawa A, Kusakizako T, Kumazaki K, Kashima A, Ishitani R, Sugita Y et al (2015) Crystal structures of SecYEG in lipidic cubic phase elucidate a precise resting and a peptide‐bound state. Cell Rep 13: 1561–1568 - PubMed
-
- Nishiyama KI, Suzuki H, Tokuda H (2000) Role of the non‐essential region encompassing the N‐terminal two transmembrane stretches of Escherichia coli SecE. Biosci Biotechnol Biochem 64: 2121–2127 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

