Strategies for Obtaining and Pruning Imputed Whole-Genome Sequence Data for Genomic Prediction
- PMID: 31379929
- PMCID: PMC6650575
- DOI: 10.3389/fgene.2019.00673
Strategies for Obtaining and Pruning Imputed Whole-Genome Sequence Data for Genomic Prediction
Abstract
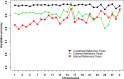
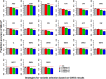
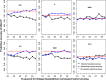
Genomic prediction with imputed whole-genome sequencing (WGS) data is an attractive approach to improve predictive ability with low cost. However, high accuracy has not been realized using this method in livestock. In this study, we imputed 435 individuals from 600K single nucleotide polymorphism (SNP) chip data to WGS data using different reference panels. We also investigated the prediction accuracy of genomic best linear unbiased prediction (GBLUP) using imputed WGS data from different reference panels, linkage disequilibrium (LD)-based marker pruning, and pre-selected variants based on Genome-wide association society (GWAS) results. Results showed that the imputation accuracies from 600K to WGS data were 0.873 ± 0.038, 0.906 ± 0.036, and 0.979 ± 0.010 for the internal, external, and combined reference panels, respectively. In most traits of chickens, the prediction accuracy of imputed WGS data obtained from the internal reference panel was greater than or equal to that of the combined reference panel; the external reference panel had the lowest prediction accuracy. Compared with 600K chip data, GBLUP with imputed WGS data had only a small increase (1-3%) in prediction accuracy. Using only variants selected from imputed WGS data based on GWAS results resulted in almost no increase for most traits and even increased the bias of the regression coefficient. The impact of the degree of LD of selected and remaining variants on prediction accuracy was different. For average daily gain (ADG), residual feed intake (RFI), intestine length (IL), and body weight in 91 days (BW91), the accuracy of GBLUP increased as the degree of LD of selected variants decreased, but the opposite relationship occurred for the remaining variants. But for breast muscle weight (BMW) and average daily feed intake (ADFI), the accuracy of GBLUP increased as the degree of LD of selected variants increased, and the degree of LD of remaining variants had a small effect on prediction accuracy. Overall, the optimal imputation strategy to obtain WGS data for genomic prediction should consider the relationship between selected individuals and target population individuals to avoid heterogeneity of imputation. LD-based marker pruning can be used to improve the accuracy of genomic prediction using imputed WGS data.
Keywords: GWAS; LD-based marker pruning; chickens; genomic prediction; imputed WGS data.
Figures

References
Associated data
LinkOut - more resources
Full Text Sources
Research Materials

