The Enigmatic Roles of PPR-SMR Proteins in Plants
- PMID: 31380188
- PMCID: PMC6662315
- DOI: 10.1002/advs.201900361
The Enigmatic Roles of PPR-SMR Proteins in Plants
Abstract
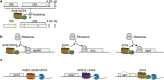
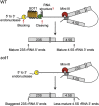
The pentatricopeptide repeat (PPR) protein family, with more than 400 members, is one of the largest and most diverse protein families in land plants. A small subset of PPR proteins contain a C-terminal small MutS-related (SMR) domain. Although there are relatively few PPR-SMR proteins, they play essential roles in embryo development, chloroplast biogenesis and gene expression, and plastid-to-nucleus retrograde signaling. Here, recent advances in understanding the roles of PPR-SMR proteins and the SMR domain based on a combination of genetic, biochemical, and physiological analyses are described. In addition, the potential of the PPR-SMR protein SOT1 to serve as a tool for RNA manipulation is highlighted.
Keywords: RNA endonuclease; RNA manipulation; pentatricopeptide repeat‐small MutSrelate (PPR‐SMR) proteins.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
PPR-SMR protein SOT1 has RNA endonuclease activity.Proc Natl Acad Sci U S A. 2017 Feb 21;114(8):E1554-E1563. doi: 10.1073/pnas.1612460114. Epub 2017 Feb 6. Proc Natl Acad Sci U S A. 2017. PMID: 28167782 Free PMC article.
-
PPR-SMRs: ancient proteins with enigmatic functions.RNA Biol. 2013;10(9):1501-10. doi: 10.4161/rna.26172. Epub 2013 Aug 28. RNA Biol. 2013. PMID: 24004908 Free PMC article. Review.
-
Moss PPR-SMR protein PpPPR_64 influences the expression of a psaA-psaB-rps14 gene cluster and processing of the 23S-4.5S rRNA precursor in chloroplasts.Plant Mol Biol. 2021 Nov;107(4-5):417-429. doi: 10.1007/s11103-020-01090-z. Epub 2020 Oct 31. Plant Mol Biol. 2021. PMID: 33128724
-
The PPR-SMR protein PPR53 enhances the stability and translation of specific chloroplast RNAs in maize.Plant J. 2016 Mar;85(5):594-606. doi: 10.1111/tpj.13093. Epub 2016 Feb 5. Plant J. 2016. PMID: 26643268 Free PMC article.
-
GUN1 and Plastid RNA Metabolism: Learning from Genetics.Cells. 2020 Oct 16;9(10):2307. doi: 10.3390/cells9102307. Cells. 2020. PMID: 33081381 Free PMC article. Review.
Cited by
-
The PPR-Domain Protein SOAR1 Regulates Salt Tolerance in Rice.Rice (N Y). 2022 Dec 3;15(1):62. doi: 10.1186/s12284-022-00608-x. Rice (N Y). 2022. PMID: 36463341 Free PMC article.
-
Organellar and Secretory Ribonucleases: Major Players in Plant RNA Homeostasis.Plant Physiol. 2020 Aug;183(4):1438-1452. doi: 10.1104/pp.20.00076. Epub 2020 Jun 8. Plant Physiol. 2020. PMID: 32513833 Free PMC article.
-
The small PPR protein SPR2 interacts with PPR-SMR1 to facilitate the splicing of introns in maize mitochondria.Plant Physiol. 2022 Oct 27;190(3):1763-1776. doi: 10.1093/plphys/kiac379. Plant Physiol. 2022. PMID: 35976145 Free PMC article.
-
The Role of Chloroplast Gene Expression in Plant Responses to Environmental Stress.Int J Mol Sci. 2020 Aug 24;21(17):6082. doi: 10.3390/ijms21176082. Int J Mol Sci. 2020. PMID: 32846932 Free PMC article. Review.
-
The PPR-SMR Protein ATP4 Is Required for Editing the Chloroplast rps8 mRNA in Rice and Maize.Plant Physiol. 2020 Dec;184(4):2011-2021. doi: 10.1104/pp.20.00849. Epub 2020 Sep 14. Plant Physiol. 2020. PMID: 32928899 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
