Convergent evolution in the mechanisms of ACBD3 recruitment to picornavirus replication sites
- PMID: 31381608
- PMCID: PMC6695192
- DOI: 10.1371/journal.ppat.1007962
Convergent evolution in the mechanisms of ACBD3 recruitment to picornavirus replication sites
Abstract
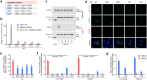
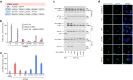
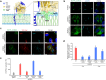
Enteroviruses, members of the family of picornaviruses, are the most common viral infectious agents in humans causing a broad spectrum of diseases ranging from mild respiratory illnesses to life-threatening infections. To efficiently replicate within the host cell, enteroviruses hijack several host factors, such as ACBD3. ACBD3 facilitates replication of various enterovirus species, however, structural determinants of ACBD3 recruitment to the viral replication sites are poorly understood. Here, we present a structural characterization of the interaction between ACBD3 and the non-structural 3A proteins of four representative enteroviruses (poliovirus, enterovirus A71, enterovirus D68, and rhinovirus B14). In addition, we describe the details of the 3A-3A interaction causing the assembly of the ACBD3-3A heterotetramers and the interaction between the ACBD3-3A complex and the lipid bilayer. Using structure-guided identification of the point mutations disrupting these interactions, we demonstrate their roles in the intracellular localization of these proteins, recruitment of downstream effectors of ACBD3, and facilitation of enterovirus replication. These structures uncovered a striking convergence in the mechanisms of how enteroviruses and kobuviruses, members of a distinct group of picornaviruses that also rely on ACBD3, recruit ACBD3 and its downstream effectors to the sites of viral replication.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Similar articles
-
ACBD3 Is an Essential Pan-enterovirus Host Factor That Mediates the Interaction between Viral 3A Protein and Cellular Protein PI4KB.mBio. 2019 Feb 12;10(1):e02742-18. doi: 10.1128/mBio.02742-18. mBio. 2019. PMID: 30755512 Free PMC article.
-
The 3A protein from multiple picornaviruses utilizes the golgi adaptor protein ACBD3 to recruit PI4KIIIβ.J Virol. 2012 Apr;86(7):3605-16. doi: 10.1128/JVI.06778-11. Epub 2012 Jan 18. J Virol. 2012. PMID: 22258260 Free PMC article.
-
Recruitment of PI4KIIIβ to coxsackievirus B3 replication organelles is independent of ACBD3, GBF1, and Arf1.J Virol. 2014 Mar;88(5):2725-36. doi: 10.1128/JVI.03650-13. Epub 2013 Dec 18. J Virol. 2014. PMID: 24352456 Free PMC article.
-
Picornavirus--host interactions to construct viral secretory membranes.Prog Mol Biol Transl Sci. 2015;129:189-212. doi: 10.1016/bs.pmbts.2014.10.007. Epub 2014 Dec 8. Prog Mol Biol Transl Sci. 2015. PMID: 25595805 Review.
-
Emerging Role for Acyl-CoA Binding Domain Containing 3 at Membrane Contact Sites During Viral Infection.Front Microbiol. 2020 Apr 8;11:608. doi: 10.3389/fmicb.2020.00608. eCollection 2020. Front Microbiol. 2020. PMID: 32322249 Free PMC article. Review.
Cited by
-
Precursors of Viral Proteases as Distinct Drug Targets.Viruses. 2021 Oct 2;13(10):1981. doi: 10.3390/v13101981. Viruses. 2021. PMID: 34696411 Free PMC article. Review.
-
Picornaviruses: A View from 3A.Viruses. 2021 Mar 11;13(3):456. doi: 10.3390/v13030456. Viruses. 2021. PMID: 33799649 Free PMC article. Review.
-
Structural insights into Acyl-coenzyme A binding domain containing 3 (ACBD3) protein hijacking by picornaviruses.Protein Sci. 2019 Dec;28(12):2073-2079. doi: 10.1002/pro.3738. Epub 2019 Oct 17. Protein Sci. 2019. PMID: 31583778 Free PMC article.
-
Enterovirus A71 antivirals: Past, present, and future.Acta Pharm Sin B. 2022 Apr;12(4):1542-1566. doi: 10.1016/j.apsb.2021.08.017. Epub 2021 Aug 20. Acta Pharm Sin B. 2022. PMID: 35847514 Free PMC article.
-
Advances in anti-EV-A71 drug development research.J Adv Res. 2024 Feb;56:137-156. doi: 10.1016/j.jare.2023.03.007. Epub 2023 Mar 30. J Adv Res. 2024. PMID: 37001813 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

