Novel Vitronectin Variations and Their Comparative Analysis in Six Porcine Breeds
- PMID: 31382414
- PMCID: PMC6719903
- DOI: 10.3390/ani9080520
Novel Vitronectin Variations and Their Comparative Analysis in Six Porcine Breeds
Abstract
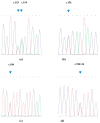
Vitronectin plays a role in the blood homeostasis and has been implicated in cell adhesion, migration, and proliferation. Vitronectin has a potential role affecting the residual feed intake (RFI) or feeding efficiency in swine production. Its variations have not been reported in Chinese swine breeds. In this study, two regions of porcine vitronectin were analyzed using PCR and sequencing. The sequence analysis revealed thirteen nucleotide substitutions in region 1 (exon 2- exon 3) and three nucleotide substitutions in region 2 (exon 5- intron 5), which would result in five amino acid changes (p.Ala52Thr, p.Leu94Pro, p.Leu94Gln, p.Gln94Pro, and p.Glu126Gly). In region 1, c.156C/T, c.281A/T, and c.377A/G were the most common (at a total frequency of 49.3%, 31.3% and 31.9% respectively), whereas c.153C/T and c.180C/G were rare (at a total frequency of 1.39%). In region 2, c.597 + 12A/G was the most common (at a total frequency of 39.6%), followed by c.597 + 15A/G (at a total frequency of 31.3%) and c.459A/G (at a total frequency of 16.0%). There was a difference (p < 0.05) in variant frequencies between Chinese breeds and overseas breeds. These results indicate that the porcine vitronectin gene is polymorphic and suggest further analysis is required to see if the variation detected affects RFI or feed efficiency in swines.
Keywords: Vitronectin; feed efficiency; pigs.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Identification of a novel premature stop codon and other recurrent variations in the porcine gelsolin gene.Gene. 2020 Sep 5;754:144879. doi: 10.1016/j.gene.2020.144879. Epub 2020 Jun 9. Gene. 2020. PMID: 32531458
-
Genetic variations of the porcine PRKAG3 gene in Chinese indigenous pig breeds.Genet Sel Evol. 2004 Jul-Aug;36(4):481-6. doi: 10.1186/1297-9686-36-4-481. Genet Sel Evol. 2004. PMID: 15231235 Free PMC article.
-
Identification of potential serum biomarkers to predict feed efficiency in young pigs.J Anim Sci. 2016 Apr;94(4):1482-92. doi: 10.2527/jas.2015-9692. J Anim Sci. 2016. PMID: 27136007
-
Detecting novel SNPs and breed-specific haplotypes at calpastatin gene in Iranian fat- and thin-tailed sheep breeds and their effects on protein structure.Gene. 2014 Mar 1;537(1):132-9. doi: 10.1016/j.gene.2013.12.023. Epub 2014 Jan 5. Gene. 2014. PMID: 24401538
-
Genetic parameters for different measures of feed efficiency and related traits in boars of three pig breeds.J Anim Sci. 2013 Sep;91(9):4069-79. doi: 10.2527/jas.2012-6197. Epub 2013 Jul 3. J Anim Sci. 2013. PMID: 23825329
References
-
- Bittorf S.V., Williams E.C., Mosher D.F. Alteration of vitronectin. Characterization of changes induced by treatment with urea. J. Biol. Chem. 1993;268:24838–24846. - PubMed
-
- Zhang D., Hudson A.E., Delostrinos C.F., Carmean N., Eastman R., Jr., Hicks B., Hurst R.E., Bassuk J.A. Dual sources of vitronectin in the human lower urinary tract: Synthesis by urothelium vs. extravasation from the bloodstream. Am. J. Physiol.-Ren. Physiol. 2011;300:475–487. doi: 10.1152/ajprenal.00407.2010. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

