Targeted Activation of Cystic Fibrosis Transmembrane Conductance Regulator
- PMID: 31383454
- PMCID: PMC6822231
- DOI: 10.1016/j.ymthe.2019.07.002
Targeted Activation of Cystic Fibrosis Transmembrane Conductance Regulator
Abstract
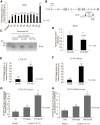
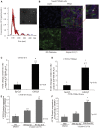
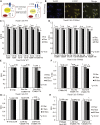
Cystic fibrosis (CF) is caused by mutations in the CF transmembrane conductance regulator (CFTR) gene. The majority of CFTR mutations result in impaired chloride channel function as only a fraction of the mutated CFTR reaches the plasma membrane. The development of a therapeutic approach that facilitates increased cell-surface expression of CFTR could prove clinically relevant. Here, we evaluate and contrast two molecular approaches to activate CFTR expression. We find that an RNA-guided nuclease null Cas9 (dCas9) fused with a tripartite activator, VP64-p65-Rta can activate endogenous CFTR in cultured human nasal epithelial cells from CF patients. We also find that targeting BGas, a long non-coding RNA involved in transcriptionally modulating CFTR expression with a gapmer, induced both strong knockdown of BGas and concordant activation of CFTR. Notably, the gapmer can be delivered to target cells when generated as electrostatic particles with recombinant HIV-Tat cell penetrating peptide (CPP), when packaged into exosomes, or when loaded into lipid nanoparticles (LNPs). Treatment of patient-derived human nasal epithelial cells containing F508del with gapmer-CPP, gapmer-exosomes, or LNPs resulted in increased expression and function of CFTR. Collectively, these observations suggest that CRISPR/dCas-VPR (CRISPR) and BGas-gapmer approaches can target and specifically activate CFTR.
Keywords: ASO delivery; CFTR; Tat-CPP; cystic fibrosis; exosome; lncRNA BGas; transcriptional regulation.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Mesenchymal Stem Cell exosome delivered Zinc Finger Protein activation of cystic fibrosis transmembrane conductance regulator.J Extracell Vesicles. 2021 Jan;10(3):e12053. doi: 10.1002/jev2.12053. Epub 2021 Jan 23. J Extracell Vesicles. 2021. PMID: 33532041 Free PMC article.
-
Long Non-coding RNA BGas Regulates the Cystic Fibrosis Transmembrane Conductance Regulator.Mol Ther. 2016 Aug;24(8):1351-7. doi: 10.1038/mt.2016.112. Epub 2016 May 30. Mol Ther. 2016. PMID: 27434588 Free PMC article.
-
The anion transporter SLC26A9 localizes to tight junctions and is degraded by the proteasome when co-expressed with F508del-CFTR.J Biol Chem. 2019 Nov 29;294(48):18269-18284. doi: 10.1074/jbc.RA119.010192. Epub 2019 Oct 23. J Biol Chem. 2019. PMID: 31645438 Free PMC article.
-
CFTR-NHERF2-LPA₂ Complex in the Airway and Gut Epithelia.Int J Mol Sci. 2017 Sep 4;18(9):1896. doi: 10.3390/ijms18091896. Int J Mol Sci. 2017. PMID: 28869532 Free PMC article. Review.
-
Repairing the basic defect in cystic fibrosis - one approach is not enough.FEBS J. 2016 Jan;283(2):246-64. doi: 10.1111/febs.13531. Epub 2015 Oct 18. FEBS J. 2016. PMID: 26416076 Review.
Cited by
-
Understanding CFTR Functionality: A Comprehensive Review of Tests and Modulator Therapy in Cystic Fibrosis.Cell Biochem Biophys. 2024 Mar;82(1):15-34. doi: 10.1007/s12013-023-01200-w. Epub 2023 Dec 4. Cell Biochem Biophys. 2024. PMID: 38048024 Review.
-
New Directions in Pulmonary Gene Therapy.Hum Gene Ther. 2020 Sep;31(17-18):921-939. doi: 10.1089/hum.2020.166. Hum Gene Ther. 2020. PMID: 32814451 Free PMC article. Review.
-
Treatment of Cystic Fibrosis: From Gene- to Cell-Based Therapies.Front Pharmacol. 2021 Mar 16;12:639475. doi: 10.3389/fphar.2021.639475. eCollection 2021. Front Pharmacol. 2021. PMID: 33796025 Free PMC article. Review.
-
Epigenome editing of the CFTR-locus for treatment of cystic fibrosis.J Cyst Fibros. 2022 Jan;21(1):164-171. doi: 10.1016/j.jcf.2021.04.008. Epub 2021 May 25. J Cyst Fibros. 2022. PMID: 34049825 Free PMC article.
-
A small molecule that induces translational readthrough of CFTR nonsense mutations by eRF1 depletion.Nat Commun. 2021 Jul 16;12(1):4358. doi: 10.1038/s41467-021-24575-x. Nat Commun. 2021. PMID: 34272367 Free PMC article.
References
-
- Kerem B., Rommens J.M., Buchanan J.A., Markiewicz D., Cox T.K., Chakravarti A., Buchwald M., Tsui L.C. Identification of the cystic fibrosis gene: genetic analysis. Science. 1989;245:1073–1080. - PubMed
-
- Sheppard D.N., Welsh M.J. Structure and function of the CFTR chloride channel. Physiol. Rev. 1999;79(1 Suppl):S23–S45. - PubMed
-
- Dalemans W., Barbry P., Champigny G., Jallat S., Dott K., Dreyer D., Crystal R.G., Pavirani A., Lecocq J.P., Lazdunski M. Altered chloride ion channel kinetics associated with the delta F508 cystic fibrosis mutation. Nature. 1991;354:526–528. - PubMed
-
- Bobadilla J.L., Macek M., Jr., Fine J.P., Farrell P.M. Cystic fibrosis: a worldwide analysis of CFTR mutations--correlation with incidence data and application to screening. Hum. Mutat. 2002;19:575–606. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

