Brain-targeted drug delivery by manipulating protein corona functions
- PMID: 31395892
- PMCID: PMC6687821
- DOI: 10.1038/s41467-019-11593-z
Brain-targeted drug delivery by manipulating protein corona functions
Abstract
Protein corona presents a major obstacle to bench-to-bedside translation of targeted drug delivery systems, severely affecting targeting yields and directing unfavorable biodistribution. Corona-mediated targeting provides a new impetus for specific drug delivery by precisely manipulating interaction modes of functional plasma proteins on nano-surface. Here bio-inspired liposomes (SP-sLip) were developed by modifying liposomal surface with a short nontoxic peptide derived from Aβ1-42 that specifically interacts with the lipid-binding domain of exchangeable apolipoproteins. SP-sLip absorb plasma apolipoproteins A1, E and J, consequently exposing receptor-binding domain of apolipoproteins to achieve brain-targeted delivery. Doxorubicin loaded SP-sLip (SP-sLip/DOX) show significant enhancement of brain distribution and anti-brain cancer effect in comparison to doxorubicin loaded plain liposomes. SP-sLip preserve functions of the absorbed human plasma ApoE, and the corona-mediated targeting strategy works in SP modified PLGA nanoparticles. The present study may pave a new avenue to facilitate clinical translation of targeted drug delivery systems.
Conflict of interest statement
C.Z. and Z.Z. are inventors on the patent related to this work that has been filed by Fudan University, Shanghai, China (PCT/CN2019/087363). The other authors declare no competing interests.
Figures





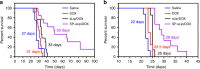


References
-
- Langer R. Drug delivery and targeting. Nature. 1998;392:5–10. - PubMed
-
- van der Meel, R., Vehmeijer, L. J. C., Kok, R. J., Storm, G. & van Gaal, E. V. B. in Intracellular Delivery III: Market Entry Barriers of Nanomedicines (eds Aleš, P. & Volkmar, W.) 163–200 (Springer International Publishing, Switzerland, 2016).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

