Biocontrol of Aspergillus flavus in Ensiled Sorghum by Water Kefir Microorganisms
- PMID: 31405185
- PMCID: PMC6722955
- DOI: 10.3390/microorganisms7080253
Biocontrol of Aspergillus flavus in Ensiled Sorghum by Water Kefir Microorganisms
Abstract
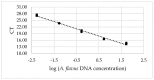
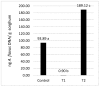
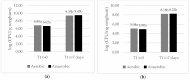
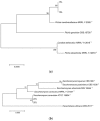
The capacity of microorganisms from water kefir (WK) to control Aspergillus flavus growth during the aerobic phase of ensiled sorghum grains was determined. Sorghum inoculated with A. flavus was treated with filter-sterilized and non-sterilized water kefir, ensiled, and incubated 7 days at 25 °C. A. flavus growth was quantified by qPCR after incubation. Mold growth was inhibited in the presence of water kefir while no inhibition was observed when filter-sterilized water kefir was applied, demonstrating the relevant role of the microorganisms in the kefir water in the biocontrol process. Fungal and bacterial diversity in treated sorghum mini-silos was analyzed by high-throughput sequencing. Firmicutes was the predominant bacterial phyla and Lactobacillus represented the most abundant genus, while Ascomycota was the predominant fungal phyla with Saccharomyces and Pichia as the major genera. Bacterial and yeast counts before and after incubation indicated that the microbial community obtained from WK was able to grow in the sorghum mini-silos in the presence of A. flavus. Results of the present work indicate that the use of a mixed inoculum of microorganisms present in WK may represent an alternative management practice to avoid the growth of A. flavus in ensiled sorghum grains and the concomitant contamination with aflatoxins.
Keywords: Aspergillus flavus; high moisture sorghum silage; high-throughput sequencing; water kefir.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


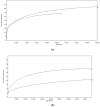
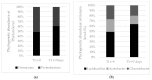


References
-
- Pahlow G., Muck R.E., Driehuis F., Oude Elferink S.J.W.H., Spoelstra S.F. Microbiology of ensiling. In: Buxton D.R., Muck R.E., Harrison J., editors. Silage Science and Technology. American Society of Agronomy; Madison, WI, USA: 2003. pp. 31–93.
-
- Scarpitta N. Qué necesitamos conocer sobre el silo de grano húmedo de sorgo?^ cNicolás Scarpitta. Rev. Plan Agropecu. 2008;126:48–54.
-
- Chiossone J., Misi O., Vicini R. [(accessed on 1 June 2019)]; Available online: http://inta.gob.ar/documentos/silaje-de-sorgo/
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

