The CTIP-mediated repair of TNF-α-induced DNA double-strand break was impaired by miR-130b in cervical cancer cell
- PMID: 31418900
- PMCID: PMC6852181
- DOI: 10.1002/cbf.3430
The CTIP-mediated repair of TNF-α-induced DNA double-strand break was impaired by miR-130b in cervical cancer cell
Abstract
Chemotherapeutic drugs that induce DNA damage have the potential to kill cancer cells, but DNA repair protects cells from damage-induced cell death. Thus, eliminating DNA repair is a potential approach to overcome cell drug resistance. In this study, we observed that the gene expression of C-terminal binding protein interacting protein (CTIP) was promoted by TNF-α stimulation and prevented TNF-α-induced double-strand breaks (DSBs) in the genomes of cervical cancer cells. The putative miR-130b targeted site within 3' untranslated region (UTR) of CTIP mRNA was identified through in silico analysis and confirmed based on experimental data. By targeting the CTIP gene, miR-130b caused the accumulation of DSBs and accelerated cell apoptosis in combination with poly ADP ribose polymerase (PARP) inhibitors. Additionally, overexpression of the CTIP gene elevated cancer cell viability by promoting proliferation while miR-130b antagonized CTIP-stimulated cell reproduction. Consequently, miR-130b destruction of DNA repair should be employed as a strategy to treat cervical cancer. SIGNIFICANCE OF THE STUDY: Cervical cancer threatens the health of women all over the world. In this study, we observed that miR-130b was able to cause the accumulation of DNA double-strand breaks through suppressing the gene expression of C-terminal binding protein interacting protein and to accelerate cell apoptosis by preventing DNA damage repairs in cervical cancer cells. As far as we know, the impact of miR-130b on the DNA double-strand break repair and on the cell apoptosis induced by the destruction of DNA repair in cervical cancer cells was firstly documented. It is reasonable to believe that miR-130b destruction of DNA repair may be employed as a strategy to treat cervical cancer in the future.
Keywords: C-terminal binding protein interacting protein; apoptosis; double-strand break; miR-130b; proliferation.
© 2019 The Authors. Cell Biochemistry and Function published by John Wiley & Sons Ltd.
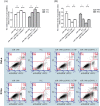
Figures



Similar articles
-
Promotion of Cervical Cancer Cell Proliferation by miR-130b Expression Level Changes and Inhibition of its Apoptosis by Targeting CDKN1A Gene.Curr Cancer Drug Targets. 2022;22(2):153-168. doi: 10.2174/1568009622666220111090715. Curr Cancer Drug Targets. 2022. PMID: 35016595 Free PMC article.
-
The TNF-α-induced expression of miR-130b protects cervical cancer cells from the cytotoxicity of TNF-α.FEBS Open Bio. 2018 Feb 16;8(4):614-627. doi: 10.1002/2211-5463.12395. eCollection 2018 Apr. FEBS Open Bio. 2018. PMID: 29632814 Free PMC article.
-
Identification of a miniature Sae2/Ctp1/CtIP ortholog from Paramecium tetraurelia required for sexual reproduction and DNA double-strand break repair.DNA Repair (Amst). 2019 May;77:96-108. doi: 10.1016/j.dnarep.2019.03.011. Epub 2019 Mar 22. DNA Repair (Amst). 2019. PMID: 30928893
-
MiR-130b/TNF-α/NF-κB/VEGFA loop inhibits prostate cancer angiogenesis.Clin Transl Oncol. 2020 Jan;22(1):111-121. doi: 10.1007/s12094-019-02217-5. Epub 2019 Oct 30. Clin Transl Oncol. 2020. PMID: 31667686
-
microRNAs: The Short Link between Cancer and RT-Induced DNA Damage Response.Front Oncol. 2014 Jun 4;4:133. doi: 10.3389/fonc.2014.00133. eCollection 2014. Front Oncol. 2014. PMID: 24926436 Free PMC article. Review. No abstract available.
Cited by
-
Home and Away: The Role of Non-Coding RNA in Intracellular and Intercellular DNA Damage Response.Genes (Basel). 2021 Sep 23;12(10):1475. doi: 10.3390/genes12101475. Genes (Basel). 2021. PMID: 34680868 Free PMC article. Review.
-
The Role of MicroRNA in DNA Damage Response.Front Genet. 2022 May 3;13:850038. doi: 10.3389/fgene.2022.850038. eCollection 2022. Front Genet. 2022. PMID: 35591858 Free PMC article. Review.
-
Multi-regulatory potency of USP1 on inflammasome components promotes pyroptosis in thyroid follicular cells and contributes to the progression of Hashimoto's thyroiditis.Mol Med. 2024 Aug 12;30(1):121. doi: 10.1186/s10020-024-00885-w. Mol Med. 2024. PMID: 39134949 Free PMC article.
-
Interactions between miRNAs and Double-Strand Breaks DNA Repair Genes, Pursuing a Fine-Tuning of Repair.Int J Mol Sci. 2022 Mar 17;23(6):3231. doi: 10.3390/ijms23063231. Int J Mol Sci. 2022. PMID: 35328651 Free PMC article. Review.
-
Jack of all trades? The versatility of RNA in DNA double-strand break repair.Essays Biochem. 2020 Oct 26;64(5):721-735. doi: 10.1042/EBC20200008. Essays Biochem. 2020. PMID: 32618336 Free PMC article. Review.

