High-throughput droplet-based microfluidics for directed evolution of enzymes
- PMID: 31433062
- PMCID: PMC6899980
- DOI: 10.1002/elps.201900222
High-throughput droplet-based microfluidics for directed evolution of enzymes
Abstract
Natural enzymes have evolved over millions of years to allow for their effective operation within specific environments. However, it is significant to note that despite their wide structural and chemical diversity, relatively few natural enzymes have been successfully applied to industrial processes. To address this limitation, directed evolution (DE) (a method that mimics the process of natural selection to evolve proteins toward a user-defined goal) coupled with droplet-based microfluidics allows the detailed analysis of millions of enzyme variants on ultra-short timescales, and thus the design of novel enzymes with bespoke properties. In this review, we aim at presenting the development of DE over the last years and highlighting the most important advancements in droplet-based microfluidics, made in this context towards the high-throughput demands of enzyme optimization. Specifically, an overview of the range of microfluidic unit operations available for the construction of DE platforms is provided, focusing on their suitability and benefits for cell-based assays, as in the case of directed evolution experimentations.
Keywords: directed evolution; droplet-based microfluidics; high throughput screening; protein engineering; single-cell.
© 2019 The Authors. Electrophoresis published by WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures

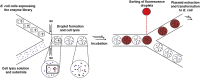

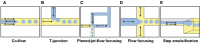

References
-
- Chapman, J. , Ismail, A. E. , Dinu, C. Z. , Catalysts 2018, 8, 238.
-
- DiCosimo, R. , McAuliffe, J. , Poulose, A. J. , Bohlmann, G. , Chem. Soc. Rev. 2013, 42, 6437–6474. - PubMed
-
- Janssen, D. B. , Dinkla, I. J. T. , Poelarends, G. J. , Terpstra, P. , Environ. Microbiol. 2005, 7, 1868–1882. - PubMed
-
- Martínez, R. , Schwaneberg, U. , Biol. Res. 2013, 46, 395–405. - PubMed
-
- Packer, M. S. , Liu, D. R. , Nat. Rev. Genet. 2015, 16, 379. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

