Population size estimation for quality control of ChIP-Seq datasets
- PMID: 31465497
- PMCID: PMC6715275
- DOI: 10.1371/journal.pone.0221760
Population size estimation for quality control of ChIP-Seq datasets
Abstract
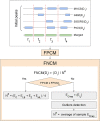
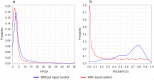
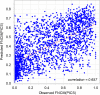
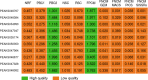
Chromatin immunoprecipitation followed by sequencing, i.e. ChIP-Seq, is a widely used experimental technology for the identification of functional protein-DNA interactions. Nowadays, such databases as ENCODE, GTRD, ChIP-Atlas and ReMap systematically collect and annotate a large number of ChIP-Seq datasets. Comprehensive control of dataset quality is currently indispensable to select the most reliable data for further analysis. In addition to existing quality control metrics, we have developed two novel metrics that allow to control false positives and false negatives in ChIP-Seq datasets. For this purpose, we have adapted well-known population size estimate for determination of unknown number of genuine transcription factor binding regions. Determination of the proposed metrics was based on overlapping distinct binding sites derived from processing one ChIP-Seq experiment by different peak callers. Moreover, the metrics also can be useful for assessing quality of datasets obtained from processing distinct ChIP-Seq experiments by a given peak caller. We also have shown that these metrics appear to be useful not only for dataset selection but also for comparison of peak callers and identification of site motifs based on ChIP-Seq datasets. The developed algorithm for determination of the false positive control metric and false negative control metric for ChIP-Seq datasets was implemented as a plugin for a BioUML platform: https://ict.biouml.org/bioumlweb/chipseq_analysis.html.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

