Gene Expression Predicts Histological Severity and Reveals Distinct Molecular Profiles of Nonalcoholic Fatty Liver Disease
- PMID: 31467298
- PMCID: PMC6715650
- DOI: 10.1038/s41598-019-48746-5
Gene Expression Predicts Histological Severity and Reveals Distinct Molecular Profiles of Nonalcoholic Fatty Liver Disease
Abstract
The heterogeneity of biological processes driving the severity of nonalcoholic fatty liver disease (NAFLD) as reflected in the transcriptome and the relationship between the pathways involved are not well established. Well-defined associations between gene expression profiles and disease progression would benefit efforts to develop novel therapies and to understand disease heterogeneity. We analyzed hepatic gene expression in controls and a cohort with the full histological spectrum of NAFLD. Protein-protein interaction and gene set variation analysis revealed distinct sets of coordinately regulated genes and pathways whose expression progressively change over the course of the disease. The progressive nature of these changes enabled us to develop a framework for calculating a disease progression score for individual genes. We show that, in aggregate, these scores correlate strongly with histological measures of disease progression and can thus themselves serve as a proxy for severity. Furthermore, we demonstrate that the expression levels of a small number of genes (~20) can be used to infer disease severity. Finally, we show that patient subgroups can be distinguished by the relative distribution of gene-level scores in specific gene sets. While future work is required to identify the specific disease characteristics that correspond to patient clusters identified on this basis, this work provides a general framework for the use of high-content molecular profiling to identify NAFLD patient subgroups.
Conflict of interest statement
Arun Sanyal: Dr. Sanyal is President of Sanyal Biotechnology and has stock options in Genfit, Akarna, Tiziana, Indalo, Durect. He has served as a consultant to Hemoshear, Echosens, AbbVie, Astra Zeneca, Nitto Denko, Ardelyx, Conatus, Nimbus, Amarin, Salix, Tobira, Takeda, Novo Nordisk, Fibrogen, Jannsen, Gilead, Boehringer, Lilly, Zafgen, Novartis, Pfizer, Immuron, Exhalenz and Genfit. He has been an unpaid consultant to Intercept, Immuron, Galectin, Fractyl, Syntlogic, Affimune, Chemomab, Nordic Bioscience and Bristol Myers Squibb. His institution has received grant support from Gilead, Salix, Tobira, Bristol Myers, Shire, Echosens, Intercept, Merck, Astra Zeneca, Malinckrodt, Cumberland and Novartis. He receives royalties from Elsevier and UptoDate. Stephen Hoang, Ryan Feaver, Banumathi Cole, Mark Lawson, Nathan Day, Justin Taylor, and Brian Wamhoff are Employees of HemoShear Therapeutics.
Figures

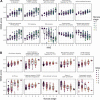




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

