Structural Insights into Curli CsgA Cross-β Fibril Architecture Inspire Repurposing of Anti-amyloid Compounds as Anti-biofilm Agents
- PMID: 31469892
- PMCID: PMC6748439
- DOI: 10.1371/journal.ppat.1007978
Structural Insights into Curli CsgA Cross-β Fibril Architecture Inspire Repurposing of Anti-amyloid Compounds as Anti-biofilm Agents
Abstract
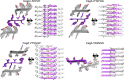
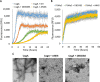
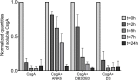
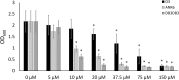
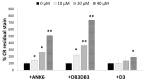
Curli amyloid fibrils secreted by Enterobacteriaceae mediate host cell adhesion and contribute to biofilm formation, thereby promoting bacterial resistance to environmental stressors. Here, we present crystal structures of amyloid-forming segments from the major curli subunit, CsgA, revealing steric zipper fibrils of tightly mated β-sheets, demonstrating a structural link between curli and human pathological amyloids. D-enantiomeric peptides, originally developed to interfere with Alzheimer's disease-associated amyloid-β, inhibited CsgA fibrillation and reduced biofilm formation in Salmonella typhimurium. Moreover, as previously shown, CsgA fibrils cross-seeded fibrillation of amyloid-β, providing support for the proposed structural resemblance and potential for cross-species amyloid interactions. The presented findings provide structural insights into amyloidogenic regions important for curli formation, suggest a novel strategy for disrupting amyloid-structured biofilms, and hypothesize on the formation of self-propagating prion-like species originating from a microbial source that could influence neurodegenerative diseases.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
-
- Dobson CM. Protein misfolding, evolution and disease. Trends Biochem Sci. 1999;24(9):329–32. . - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

