Comprehensive genomic profiling of glioblastoma tumors, BTICs, and xenografts reveals stability and adaptation to growth environments
- PMID: 31471491
- PMCID: PMC6754609
- DOI: 10.1073/pnas.1813495116
Comprehensive genomic profiling of glioblastoma tumors, BTICs, and xenografts reveals stability and adaptation to growth environments
Abstract
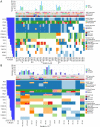
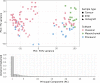
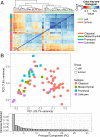
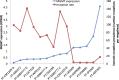
Glioblastoma multiforme (GBM) is the most deadly brain tumor, and currently lacks effective treatment options. Brain tumor-initiating cells (BTICs) and orthotopic xenografts are widely used in investigating GBM biology and new therapies for this aggressive disease. However, the genomic characteristics and molecular resemblance of these models to GBM tumors remain undetermined. We used massively parallel sequencing technology to decode the genomes and transcriptomes of BTICs and xenografts and their matched tumors in order to delineate the potential impacts of the distinct growth environments. Using data generated from whole-genome sequencing of 201 samples and RNA sequencing of 118 samples, we show that BTICs and xenografts resemble their parental tumor at the genomic level but differ at the mRNA expression and epigenomic levels, likely due to the different growth environment for each sample type. These findings suggest that a comprehensive genomic understanding of in vitro and in vivo GBM model systems is crucial for interpreting data from drug screens, and can help control for biases introduced by cell-culture conditions and the microenvironment in mouse models. We also found that lack of MGMT expression in pretreated GBM is linked to hypermutation, which in turn contributes to increased genomic heterogeneity and requires new strategies for GBM treatment.
Keywords: BTICs; genome; glioblastoma; therapy; transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Bmi1 regulates human glioblastoma stem cells through activation of differential gene networks in CD133+ brain tumor initiating cells.J Neurooncol. 2019 Jul;143(3):417-428. doi: 10.1007/s11060-019-03192-1. Epub 2019 May 21. J Neurooncol. 2019. PMID: 31115870
-
On-target JAK2/STAT3 inhibition slows disease progression in orthotopic xenografts of human glioblastoma brain tumor stem cells.Neuro Oncol. 2013 Feb;15(2):198-207. doi: 10.1093/neuonc/nos302. Epub 2012 Dec 21. Neuro Oncol. 2013. PMID: 23262510 Free PMC article.
-
Fibrinogen in the glioblastoma microenvironment contributes to the invasiveness of brain tumor-initiating cells.Brain Pathol. 2021 Sep;31(5):e12947. doi: 10.1111/bpa.12947. Epub 2021 Mar 10. Brain Pathol. 2021. PMID: 33694259 Free PMC article.
-
"Zooming in" on Glioblastoma: Understanding Tumor Heterogeneity and its Clinical Implications in the Era of Single-Cell Ribonucleic Acid Sequencing.Neurosurgery. 2021 Feb 16;88(3):477-486. doi: 10.1093/neuros/nyaa305. Neurosurgery. 2021. PMID: 32674143 Review.
-
Comparison of glioblastoma (GBM) molecular classification methods.Semin Cancer Biol. 2018 Dec;53:201-211. doi: 10.1016/j.semcancer.2018.07.006. Epub 2018 Jul 19. Semin Cancer Biol. 2018. PMID: 30031763 Review.
Cited by
-
BI-907828, a novel potent MDM2 inhibitor, inhibits glioblastoma brain tumor stem cells in vitro and prolongs survival in orthotopic xenograft mouse models.Neuro Oncol. 2023 May 4;25(5):913-926. doi: 10.1093/neuonc/noac271. Neuro Oncol. 2023. PMID: 36521007 Free PMC article.
-
Cellular heterogeneity and therapeutic response profiling of human IDH+ glioma stem cell cultures.bioRxiv [Preprint]. 2025 Aug 1:2025.07.29.667532. doi: 10.1101/2025.07.29.667532. bioRxiv. 2025. PMID: 40766425 Free PMC article. Preprint.
-
PAM (PIK3/AKT/mTOR) signaling in glia: potential contributions to brain tumors in aging.Aging (Albany NY). 2021 Jan 5;13(1):1510-1527. doi: 10.18632/aging.202459. Epub 2021 Jan 5. Aging (Albany NY). 2021. PMID: 33472174 Free PMC article. Review.
-
Epigenetic and molecular coordination between HDAC2 and SMAD3-SKI regulates essential brain tumour stem cell characteristics.Nat Commun. 2023 Aug 19;14(1):5051. doi: 10.1038/s41467-023-40776-y. Nat Commun. 2023. PMID: 37598220 Free PMC article.
-
Patient-derived glioblastoma cell lines with conserved genome profiles of the original tissue.Sci Data. 2023 Jul 12;10(1):448. doi: 10.1038/s41597-023-02365-y. Sci Data. 2023. PMID: 37438387 Free PMC article.
References
-
- Kelly J. J., et al. , Proliferation of human glioblastoma stem cells occurs independently of exogenous mitogens. Stem Cells 27, 1722–1733 (2009). - PubMed
-
- Lee H. W., Lee K., Kim D. G., Yang H., Nam D. H., Facilitating tailored therapeutic strategies for glioblastoma through an orthotopic patient-derived xenograft platform. Histol. Histopathol. 31, 269–283 (2016). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

