Dcp2: an mRNA decapping enzyme that adopts many different shapes and forms
- PMID: 31473440
- PMCID: PMC6900585
- DOI: 10.1016/j.sbi.2019.07.009
Dcp2: an mRNA decapping enzyme that adopts many different shapes and forms
Abstract
Eukaryotic mRNAs contain a 5' cap structure that protects the transcript against rapid exonucleolytic degradation. The regulation of cellular mRNA levels therefore depends on a precise control of the mRNA decapping pathways. The major mRNA decapping enzyme in eukaryotic cells is Dcp2. It is regulated by interactions with several activators, including Dcp1, Edc1, and Edc3, as well as by an autoinhibition mechanism. The structural and mechanistical characterization of Dcp2 complexes has long been impeded by the high flexibility and dynamic nature of the enzyme. Here we review recent insights into the catalytically active conformation of the mRNA decapping complex, the mode of action of decapping activators and the large interactions network that Dcp2 is embedded in.
Copyright © 2019 The Authors. Published by Elsevier Ltd.. All rights reserved.
Figures



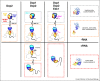

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

