Genome-wide identification of Pseudomonas syringae genes required for fitness during colonization of the leaf surface and apoplast
- PMID: 31484768
- PMCID: PMC6754560
- DOI: 10.1073/pnas.1908858116
Genome-wide identification of Pseudomonas syringae genes required for fitness during colonization of the leaf surface and apoplast
Abstract
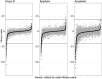
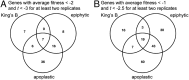
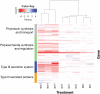
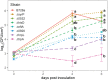
The foliar plant pathogen Pseudomonas syringae can establish large epiphytic populations on leaf surfaces before apoplastic colonization. However, the bacterial genes that contribute to these lifestyles have not been completely defined. The fitness contributions of 4,296 genes in P. syringae pv. syringae B728a were determined by genome-wide fitness profiling with a randomly barcoded transposon mutant library that was grown on the leaf surface and in the apoplast of the susceptible plant Phaseolus vulgaris Genes within the functional categories of amino acid and polysaccharide (including alginate) biosynthesis contributed most to fitness both on the leaf surface (epiphytic) and in the leaf interior (apoplast), while genes involved in type III secretion system and syringomycin synthesis were primarily important in the apoplast. Numerous other genes that had not been previously associated with in planta growth were also required for maximum epiphytic or apoplastic fitness. Fourteen hypothetical proteins and uncategorized glycosyltransferases were also required for maximum competitive fitness in and on leaves. For most genes, no relationship was seen between fitness in planta and either the magnitude of their expression in planta or degree of induction in planta compared to in vitro conditions measured in other studies. A lack of association of gene expression and fitness has important implications for the interpretation of transcriptional information and our broad understanding of plant-microbe interactions.
Keywords: endophytes; epiphytes; gene expression.
Conflict of interest statement
The authors declare no conflict of interest.
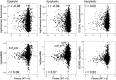
Figures
Similar articles
-
Transcriptional analysis of the global regulatory networks active in Pseudomonas syringae during leaf colonization.mBio. 2014 Sep 2;5(5):e01683-14. doi: 10.1128/mBio.01683-14. mBio. 2014. PMID: 25182327 Free PMC article.
-
Transcriptional responses of Pseudomonas syringae to growth in epiphytic versus apoplastic leaf sites.Proc Natl Acad Sci U S A. 2013 Jan 29;110(5):E425-34. doi: 10.1073/pnas.1221892110. Epub 2013 Jan 14. Proc Natl Acad Sci U S A. 2013. PMID: 23319638 Free PMC article.
-
The pH of the leaf apoplast is critical for the formation of Pseudomonas syringae-induced lesions on leaves of the common bean (Phaseolus vulgaris).Plant Sci. 2020 Jan;290:110328. doi: 10.1016/j.plantsci.2019.110328. Epub 2019 Nov 5. Plant Sci. 2020. PMID: 31779895
-
Roadmap to new virulence determinants in Pseudomonas syringae: insights from comparative genomics and genome organization.Mol Plant Microbe Interact. 2008 Jun;21(6):685-700. doi: 10.1094/MPMI-21-6-0685. Mol Plant Microbe Interact. 2008. PMID: 18624633 Review.
-
Closing the circle on the discovery of genes encoding Hrp regulon members and type III secretion system effectors in the genomes of three model Pseudomonas syringae strains.Mol Plant Microbe Interact. 2006 Nov;19(11):1151-8. doi: 10.1094/MPMI-19-1151. Mol Plant Microbe Interact. 2006. PMID: 17073298 Review.
Cited by
-
The anti-sigma factor MucA is required for viability in Pseudomonas aeruginosa.Mol Microbiol. 2021 Aug;116(2):550-563. doi: 10.1111/mmi.14732. Epub 2021 May 18. Mol Microbiol. 2021. PMID: 33905139 Free PMC article.
-
Defeated by the nines: nine extracellular strategies to avoid microbe-associated molecular patterns recognition in plants.Plant Cell. 2021 Aug 13;33(7):2116-2130. doi: 10.1093/plcell/koab109. Plant Cell. 2021. PMID: 33871653 Free PMC article.
-
Phosphoribosylpyrophosphate synthetase as a metabolic valve advances Methylobacterium/Methylorubrum phyllosphere colonization and plant growth.Nat Commun. 2024 Jul 16;15(1):5969. doi: 10.1038/s41467-024-50342-9. Nat Commun. 2024. PMID: 39013920 Free PMC article.
-
Barcoded overexpression screens in gut Bacteroidales identify genes with roles in carbon utilization and stress resistance.Nat Commun. 2024 Aug 5;15(1):6618. doi: 10.1038/s41467-024-50124-3. Nat Commun. 2024. PMID: 39103350 Free PMC article.
-
Widespread horizontal gene transfer between plants and bacteria.ISME Commun. 2024 May 13;4(1):ycae073. doi: 10.1093/ismeco/ycae073. eCollection 2024 Jan. ISME Commun. 2024. PMID: 38808121 Free PMC article.
References
-
- Lindemann J., Arny D. C., Upper C. D., Use of an apparent infection threshold population of Pseudomonas syringae to predict incidence and severity of brown spot of bean. Phytopathology 74, 1334–1339 (1984).
-
- Hirano S. S., Upper C. D., Ecology and epidemiology of foliar bacterial plant pathogens. Annu. Rev. Phytopathol. 21, 243–269 (1983).
-
- van der Wal A., Leveau J. H. J., Modelling sugar diffusion across plant leaf cuticles: The effect of free water on substrate availability to phyllosphere bacteria. Environ. Microbiol. 13, 792–797 (2011). - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

