Systematic Identification of Characteristic Genes of Ovarian Clear Cell Carcinoma Compared with High-Grade Serous Carcinoma Based on RNA-Sequencing
- PMID: 31487856
- PMCID: PMC6770582
- DOI: 10.3390/ijms20184330
Systematic Identification of Characteristic Genes of Ovarian Clear Cell Carcinoma Compared with High-Grade Serous Carcinoma Based on RNA-Sequencing
Abstract
Objective: Ovarian cancer has the highest mortality among gynecological cancers. High-grade serous carcinoma (HGSC) is the most common histotype of ovarian cancer regardless of ethnicity, whereas clear cell carcinoma (CCC) is more common in East Asians than Caucasians. The elucidation of predominant signaling pathways in these cancers is the first step towards understanding their molecular mechanisms and developing their clinical management.
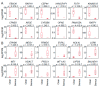
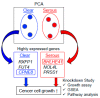
Methods: RNA sequencing was performed for 27 clinical ovarian specimens from Japanese women. Principal component analysis (PCA) was conducted on the sequence data mapped on RefSeq with normalized read counts, and functional annotation analysis was performed on genes with substantial weights in PCA. Knockdown experiments were conducted on the selected genes on the basis of PCA.
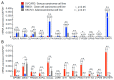
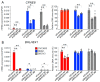
Results: Functional annotation analysis of PCA-defined genes showed predominant pathways, such as cell growth regulators and blood coagulators in CCC and transcription regulators in HGSC. Knockdown experiments showed that the inhibition of the calcium-dependent protein copine 8 (CPNE8) and the transcription factor basic helix-loop-helix family member e 41 (BHLHE41) repressed the proliferation of CCC- and HGSC-derived cells, respectively.
Conclusions: This study identified CPNE8 and BHLHE41 as characteristic genes for CCC and HGSC, respectively. The systemic identification of differentially expressed genes in CCC and HGSC will provide useful information to understand transcriptomic differences in these ovarian cancers and to further develop potential diagnostic and therapeutic options for advanced disease.
Keywords: RNA sequencing; clear cell carcinoma; gene expression; high-grade serous carcinoma; ovarian cancer.
Conflict of interest statement
The authors have no conflict of interest.
Figures


Similar articles
-
Profile of differentially expressed miRNAs in high-grade serous carcinoma and clear cell ovarian carcinoma, and the expression of miR-510 in ovarian carcinoma.Mol Med Rep. 2015 Dec;12(6):8021-31. doi: 10.3892/mmr.2015.4485. Epub 2015 Oct 26. Mol Med Rep. 2015. PMID: 26497752 Free PMC article.
-
Global miRNA expression analysis of serous and clear cell ovarian carcinomas identifies differentially expressed miRNAs including miR-200c-3p as a prognostic marker.BMC Cancer. 2014 Feb 11;14:80. doi: 10.1186/1471-2407-14-80. BMC Cancer. 2014. PMID: 24512620 Free PMC article.
-
SLFN11 is a BRCA Independent Biomarker for the Response to Platinum-Based Chemotherapy in High-Grade Serous Ovarian Cancer and Clear Cell Ovarian Carcinoma.Mol Cancer Ther. 2024 Jan 3;23(1):106-116. doi: 10.1158/1535-7163.MCT-23-0257. Mol Cancer Ther. 2024. PMID: 37717249
-
[Oncopathological aspects of BRCA1 and BRCA2 genes inactivation in tumors of ovary, fallopian tube and pelvic peritoneum].Cesk Patol. 2016 Fall;52(4):199-204. Cesk Patol. 2016. PMID: 27869446 Review. Czech.
-
High-grade serous carcinoma of tubo-ovarian origin: recent developments.Histopathology. 2017 Sep;71(3):339-356. doi: 10.1111/his.13248. Epub 2017 Jul 18. Histopathology. 2017. PMID: 28477361 Review.
Cited by
-
High-Throughput Drug Screening of Clear Cell Ovarian Cancer Organoids Reveals Vulnerability to Proteasome Inhibitors and Dinaciclib and Identifies AGR2 as a Therapeutic Target.Cancer Res Commun. 2025 Jun 1;5(6):1018-1033. doi: 10.1158/2767-9764.CRC-25-0024. Cancer Res Commun. 2025. PMID: 40459063 Free PMC article.
-
Multicellular Ovarian Cancer Model for Evaluation of Nanovector Delivery in Ascites and Metastatic Environments.Pharmaceutics. 2021 Nov 8;13(11):1891. doi: 10.3390/pharmaceutics13111891. Pharmaceutics. 2021. PMID: 34834307 Free PMC article.
-
The Role of Omics Approaches to Characterize Molecular Mechanisms of Rare Ovarian Cancers: Recent Advances and Future Perspectives.Biomedicines. 2021 Oct 15;9(10):1481. doi: 10.3390/biomedicines9101481. Biomedicines. 2021. PMID: 34680597 Free PMC article. Review.
-
Copine C plays a role in adhesion and streaming in Dictyostelium.Cell Adh Migr. 2024 Dec;18(1):1-19. doi: 10.1080/19336918.2024.2315629. Epub 2024 Feb 20. Cell Adh Migr. 2024. PMID: 38378453 Free PMC article.
-
Long Intergenic Noncoding RNA OIN1 Promotes Ovarian Cancer Growth by Modulating Apoptosis-Related Gene Expression.Int J Mol Sci. 2021 Oct 18;22(20):11242. doi: 10.3390/ijms222011242. Int J Mol Sci. 2021. PMID: 34681900 Free PMC article.
References
-
- Center for Cancer Control and Information Services, National Cancer Canter, Japan Projected Cancer Statistics, 2018. [(accessed on 15 September 2019)]; Available online: https://ganjoho.jp/en/public/statistics/short_pred.html.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

