Differential transcriptome analysis of enterohemorrhagic Escherichia coli strains reveals differences in response to plant-derived compounds
- PMID: 31488056
- PMCID: PMC6729007
- DOI: 10.1186/s12866-019-1578-4
Differential transcriptome analysis of enterohemorrhagic Escherichia coli strains reveals differences in response to plant-derived compounds
Abstract
Background: Several serious vegetable-associated outbreaks of enterohemorrhagic Escherichia coli (EHEC) infections have occurred during the last decades. In this context, vegetables have been suggested to function as secondary reservoirs for EHEC strains. Increased knowledge about the interaction of EHEC with plants including gene expression patterns in response to plant-derived compounds is required. In the current study, EHEC O157:H7 strain Sakai, EHEC O157:H- strain 3072/96, and the EHEC/enteroaggregative E. coli (EAEC) hybrid O104:H4 strain C227-11φcu were grown in lamb's lettuce medium and in M9 minimal medium to study the differential transcriptional response of these strains to plant-derived compounds with RNA-Seq technology.
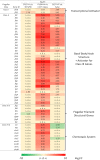
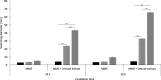
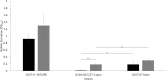
Results: Many genes involved in carbohydrate degradation and peptide utilization were similarly upregulated in all three strains, suggesting that the lamb's lettuce medium provides sufficient nutrients for proliferation. In particular, the genes galET and rbsAC involved in galactose metabolism and D-ribose catabolism, respectively, were uniformly upregulated in the investigated strains. The most prominent differences in shared genome transcript levels were observed for genes involved in the expression of flagella. Transcripts of all three classes of the flagellar hierarchy were highly abundant in strain C227-11φcu. Strain Sakai expressed only genes encoding the basal flagellar structure. In addition, both strains showed increased motility in presence of lamb's lettuce extract. Moreover, strain 3072/96 showed increased transcription activity for genes encoding the type III secretion system (T3SS) including effectors, and was identified as a powerful biofilm-producer in M9 minimal medium.
Conclusion: The current study provides clear evidence that EHEC and EHEC/EAEC strains are able to adjust their gene expression patterns towards metabolization of plant-derived compounds, demonstrating that they may proliferate well in a plant-associated environment. Moreover, we propose that flagella and other surface structures play a fundamental role in the interaction of EHEC and EHEC/EAEC with plants.
Keywords: Biofilm; Differential gene expression; EHEC; Energy metabolism; Flagella; Lettuce medium; Motility; O104:H4 strain C227–11φcu; O157:H7 strain Sakai; O157:H− strain 3072/96; Plant extract; Transcriptome.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

