Inverse pharmacology: Approaches and tools for introducing druggability into engineered proteins
- PMID: 31494210
- PMCID: PMC6891246
- DOI: 10.1016/j.biotechadv.2019.107439
Inverse pharmacology: Approaches and tools for introducing druggability into engineered proteins
Abstract
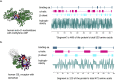
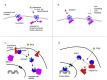
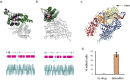
A major feature of twenty-first century medical research is the development of therapeutic strategies that use 'biologics' (large molecules, usually engineered proteins) and living cells instead of, or as well as, the small molecules that were the basis of pharmacology in earlier eras. The high power of these techniques can bring correspondingly high risk, and therefore the need for the potential for external control. One way of exerting control on therapeutic proteins is to make them responsive to small molecules; in a clinical context, these small molecules themselves have to be safe. Conventional pharmacology has resulted in thousands of small molecules licensed for use in humans, and detailed structural data on their binding to their protein targets. In principle, these data can be used to facilitate the engineering of drug-responsive modules, taken from natural proteins, into synthetic proteins. This has been done for some years (for example, Cre-ERT2) but usually in a painstaking manner. Recently, we have developed the bioinformatic tool SynPharm to facilitate the design of drug-responsive proteins. In this review, we outline the history of the field, the design and use of the Synpharm tool, and describe our own experiences in engineering druggability into the Cpf1 effector of CRISPR gene editing.
Keywords: CRISPR; Cpf1; Drug; Gene editing; Pharmaceutical; Protein engineering; Structure-function; Synthetic biology.
Copyright © 2019 The Authors. Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
SynPharm and the guide to pharmacology database: A toolset for conferring drug control on engineered proteins.Protein Sci. 2021 Jan;30(1):160-167. doi: 10.1002/pro.3971. Epub 2020 Nov 2. Protein Sci. 2021. PMID: 33047381 Free PMC article.
-
[The new generation tool for CRISPR genome editing: CRISPR/Cpf1].Sheng Wu Gong Cheng Xue Bao. 2017 Mar 25;33(3):361-371. doi: 10.13345/j.cjb.170029. Sheng Wu Gong Cheng Xue Bao. 2017. PMID: 28941336 Review. Chinese.
-
Small molecules promote CRISPR-Cpf1-mediated genome editing in human pluripotent stem cells.Nat Commun. 2018 Apr 3;9(1):1303. doi: 10.1038/s41467-018-03760-5. Nat Commun. 2018. PMID: 29610531 Free PMC article.
-
Design and assessment of engineered CRISPR-Cpf1 and its use for genome editing.Nat Protoc. 2018 May;13(5):899-914. doi: 10.1038/nprot.2018.004. Epub 2018 Apr 5. Nat Protoc. 2018. PMID: 29622802 Free PMC article.
-
The Conspicuity of CRISPR-Cpf1 System as a Significant Breakthrough in Genome Editing.Curr Microbiol. 2018 Jan;75(1):107-115. doi: 10.1007/s00284-017-1406-8. Epub 2017 Nov 30. Curr Microbiol. 2018. PMID: 29189942 Review.
Cited by
-
The IUPHAR/BPS Guide to PHARMACOLOGY in 2020: extending immunopharmacology content and introducing the IUPHAR/MMV Guide to MALARIA PHARMACOLOGY.Nucleic Acids Res. 2020 Jan 8;48(D1):D1006-D1021. doi: 10.1093/nar/gkz951. Nucleic Acids Res. 2020. PMID: 31691834 Free PMC article.
-
Accelerating therapeutic protein design with computational approaches toward the clinical stage.Comput Struct Biotechnol J. 2023 Apr 29;21:2909-2926. doi: 10.1016/j.csbj.2023.04.027. eCollection 2023. Comput Struct Biotechnol J. 2023. PMID: 38213894 Free PMC article. Review.
References
-
- Baron U., Bujard H. Tet repressor-based system for regulated gene expression in eukaryotic cells: principles and advances. Methods Enzymol. 2000;327:401–421. - PubMed
-
- Bernstein F.C., Koetzle T.F., Williams G.J., Meyer E.F., Jr., Brice M.D., Rodgers J.R., Kennard O., Shimanouchi T., Tasumi M. The protein data bank: a computer-based archival file for macromolecular structures. J. Mol. Biol. 1977;112:535–542. - PubMed
-
- Chen X., Liu M., Gilson M.K. BindingDB: a web-accessible molecular recognition database. Comb. Chem. High Throughput Screen. 2001;2001:719–725. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

