A comprehensive analysis of the B3 superfamily identifies tissue-specific and stress-responsive genes in chickpea (Cicer arietinum L.)
- PMID: 31497464
- PMCID: PMC6710316
- DOI: 10.1007/s13205-019-1875-5
A comprehensive analysis of the B3 superfamily identifies tissue-specific and stress-responsive genes in chickpea (Cicer arietinum L.)
Abstract
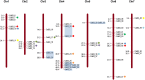
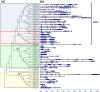
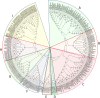
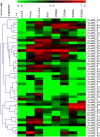
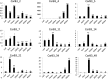
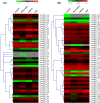
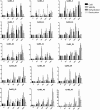
The aim of this study was to provide a comprehensive analysis of the plant-specific B3 domain-containing transcription factors (TFs) in chickpea. Scanning of the chickpea genome resulted in the identification of 51 B3 domain-containing TFs that were located on seven out of eight chickpea chromosomes. Based on the presence of additional domains other than the B3 domain, the candidates were classified into four subfamilies, i.e., ARF (24), REM (19), LAV (6) and RAV (2). Phylogenetic analysis classified them into four groups in which members of the same group had similar intron-exon organization and motif composition. Genome duplication analysis of the candidate B3 genes revealed an event of segmental duplication that was instrumental in the expansion of the B3 gene family. Ka/Ks analysis showed that the B3 gene family was under purifying selection. Further, chickpea B3 genes showed maximum orthology with Medicago followed by soybean and Arabidopsis. Promoter analyses of the B3 genes led to the identification of several tissue-specific and stress-responsive cis-regulatory elements. Expression profiling of the candidate B3 genes using publicly available RNA-seq data of several chickpea tissues indicated their putative role in plant development and abiotic stress response. These findings were further validated by real-time expression analysis. Overall, this study provides a comprehensive analysis of the B3 domain-containing proteins in chickpea that would aid in devising strategies for crop manipulation in chickpea.
Keywords: B3 domain-containing protein; Chickpea; Expression analysis; Phylogeny.
Conflict of interest statement
Conflict of interestOn behalf of all authors, the corresponding author states that there is no conflict of interest.
Figures
References
LinkOut - more resources
Full Text Sources

