A protein architecture guided screen for modification dependent restriction endonucleases
- PMID: 31504772
- PMCID: PMC6765204
- DOI: 10.1093/nar/gkz755
A protein architecture guided screen for modification dependent restriction endonucleases
Abstract
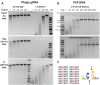
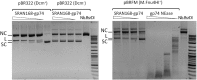
Modification dependent restriction endonucleases (MDREs) often have separate catalytic and modification dependent domains. We systematically looked for previously uncharacterized fusion proteins featuring a PUA or DUF3427 domain and HNH or PD-(D/E)XK catalytic domain. The enzymes were clustered by similarity of their putative modification sensing domains into several groups. The TspA15I (VcaM4I, CmeDI), ScoA3IV (MsiJI, VcaCI) and YenY4I groups, all featuring a PUA superfamily domain, preferentially cleaved DNA containing 5-methylcytosine or 5-hydroxymethylcytosine. ScoA3V, also featuring a PUA superfamily domain, but of a different clade, exhibited 6-methyladenine stimulated nicking activity. With few exceptions, ORFs for PUA-superfamily domain containing endonucleases were not close to DNA methyltransferase ORFs, strongly supporting modification dependent activity of the endonucleases. DUF3427 domain containing fusion proteins had very little or no endonuclease activity, despite the presence of a putative PD-(D/E)XK catalytic domain. However, their expression potently restricted phage T4gt in Escherichia coli cells. In contrast to the ORFs for PUA domain containing endonucleases, the ORFs for DUF3427 fusion proteins were frequently found in defense islands, often also featuring DNA methyltransferases.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures







Similar articles
-
Crystal structures of the EVE-HNH endonuclease VcaM4I in the presence and absence of DNA.Nucleic Acids Res. 2021 Feb 22;49(3):1708-1723. doi: 10.1093/nar/gkaa1218. Nucleic Acids Res. 2021. PMID: 33450012 Free PMC article.
-
Cloning and analysis of a bifunctional methyltransferase/restriction endonuclease TspGWI, the prototype of a Thermus sp. enzyme family.BMC Mol Biol. 2009 May 29;10:52. doi: 10.1186/1471-2199-10-52. BMC Mol Biol. 2009. PMID: 19480701 Free PMC article.
-
Characterization of AloI, a restriction-modification system of a new type.J Mol Biol. 2001 Nov 23;314(2):205-16. doi: 10.1006/jmbi.2001.5049. J Mol Biol. 2001. PMID: 11718555
-
Specificity of restriction endonucleases and DNA modification methyltransferases a review (Edition 3).Gene. 1990 Aug 16;92(1-2):1-248. doi: 10.1016/0378-1119(90)90486-b. Gene. 1990. PMID: 2172084 Review.
-
Nicking endonucleases.Biochemistry (Mosc). 2009 Dec;74(13):1457-66. doi: 10.1134/s0006297909130033. Biochemistry (Mosc). 2009. PMID: 20210703 Review.
Cited by
-
Structural analysis of the BisI family of modification dependent restriction endonucleases.Nucleic Acids Res. 2024 Aug 27;52(15):9103-9118. doi: 10.1093/nar/gkae634. Nucleic Acids Res. 2024. PMID: 39041409 Free PMC article.
-
Improving Mobilization of Foreign DNA into Zymomonas mobilis Strain ZM4 by Removal of Multiple Restriction Systems.Appl Environ Microbiol. 2021 Sep 10;87(19):e0080821. doi: 10.1128/AEM.00808-21. Epub 2021 Sep 10. Appl Environ Microbiol. 2021. PMID: 34288704 Free PMC article.
-
Crystal structures of the EVE-HNH endonuclease VcaM4I in the presence and absence of DNA.Nucleic Acids Res. 2021 Feb 22;49(3):1708-1723. doi: 10.1093/nar/gkaa1218. Nucleic Acids Res. 2021. PMID: 33450012 Free PMC article.
-
The DNA cytosine methylome revealed two methylation motifs in the upstream regions of genes related to morphological and physiological differentiation in Streptomyces coelicolor A(3)2 M145.Sci Rep. 2023 Apr 29;13(1):7038. doi: 10.1038/s41598-023-34075-1. Sci Rep. 2023. PMID: 37120673 Free PMC article.
-
Protein Domain Guided Screen for Sequence Specific and Phosphorothioate-Dependent Restriction Endonucleases.Front Microbiol. 2020 Aug 18;11:1960. doi: 10.3389/fmicb.2020.01960. eCollection 2020. Front Microbiol. 2020. PMID: 33013736 Free PMC article.
References
-
- Geier G.E., Modrich P.. Recognition sequence of the dam methylase of Escherichia coli K12 and mode of cleavage of Dpn I endonuclease. J. Biol. Chem. 1979; 254:1408–1413. - PubMed
-
- Sutherland E., Coe L., Raleigh E.A.. McrBC: a multisubunit GTP-dependent restriction endonuclease. J Mol. Biol. 1992; 225:327–348. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

