LYAR potentiates rRNA synthesis by recruiting BRD2/4 and the MYST-type acetyltransferase KAT7 to rDNA
- PMID: 31504794
- PMCID: PMC6821171
- DOI: 10.1093/nar/gkz747
LYAR potentiates rRNA synthesis by recruiting BRD2/4 and the MYST-type acetyltransferase KAT7 to rDNA
Abstract
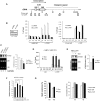
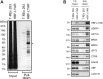
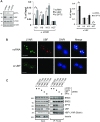
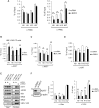
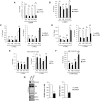
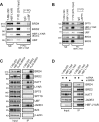
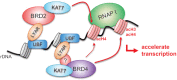
Activation of ribosomal RNA (rRNA) synthesis is pivotal during cell growth and proliferation, but its aberrant upregulation may promote tumorigenesis. Here, we demonstrate that the candidate oncoprotein, LYAR, enhances ribosomal DNA (rDNA) transcription. Our data reveal that LYAR binds the histone-associated protein BRD2 without involvement of acetyl-lysine-binding bromodomains and recruits BRD2 to the rDNA promoter and transcribed regions via association with upstream binding factor. We show that BRD2 is required for the recruitment of the MYST-type acetyltransferase KAT7 to rDNA loci, resulting in enhanced local acetylation of histone H4. In addition, LYAR binds a complex of BRD4 and KAT7, which is then recruited to rDNA independently of the BRD2-KAT7 complex to accelerate the local acetylation of both H4 and H3. BRD2 also helps recruit BRD4 to rDNA. By contrast, LYAR has no effect on rDNA methylation or the binding of RNA polymerase I subunits to rDNA. These data suggest that LYAR promotes the association of the BRD2-KAT7 and BRD4-KAT7 complexes with transcription-competent rDNA loci but not to transcriptionally silent rDNA loci, thereby increasing rRNA synthesis by altering the local acetylation status of histone H3 and H4.
© The Author(s) 2019. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

Similar articles
-
Histone acetyltransferase 7 (KAT7)-dependent intragenic histone acetylation regulates endothelial cell gene regulation.J Biol Chem. 2018 Mar 23;293(12):4381-4402. doi: 10.1074/jbc.RA117.001383. Epub 2018 Feb 6. J Biol Chem. 2018. PMID: 29414790 Free PMC article.
-
The silencing complex SAS-I links histone acetylation to the assembly of repressed chromatin by CAF-I and Asf1 in Saccharomyces cerevisiae.Genes Dev. 2001 Dec 1;15(23):3169-82. doi: 10.1101/gad.929001. Genes Dev. 2001. PMID: 11731480 Free PMC article.
-
Recruitment of the nucleolar remodeling complex NoRC establishes ribosomal DNA silencing in chromatin.Mol Cell Biol. 2004 Feb;24(4):1791-8. doi: 10.1128/MCB.24.4.1791-1798.2004. Mol Cell Biol. 2004. PMID: 14749393 Free PMC article.
-
Deciphering structure, function and mechanism of lysine acetyltransferase HBO1 in protein acetylation, transcription regulation, DNA replication and its oncogenic properties in cancer.Cell Mol Life Sci. 2020 Feb;77(4):637-649. doi: 10.1007/s00018-019-03296-x. Epub 2019 Sep 18. Cell Mol Life Sci. 2020. PMID: 31535175 Free PMC article. Review.
-
MYST-family histone acetyltransferases: beyond chromatin.Cell Mol Life Sci. 2011 Apr;68(7):1147-56. doi: 10.1007/s00018-010-0599-9. Epub 2010 Dec 4. Cell Mol Life Sci. 2011. PMID: 21132344 Free PMC article. Review.
Cited by
-
Current insights into the oncogenic roles of lncRNA LINC00355.Cancer Innov. 2023 Sep 14;2(6):448-462. doi: 10.1002/cai2.91. eCollection 2023 Dec. Cancer Innov. 2023. PMID: 38125763 Free PMC article. Review.
-
The Class IIA Histone Deacetylase (HDAC) Inhibitor TMP269 Downregulates Ribosomal Proteins and Has Anti-Proliferative and Pro-Apoptotic Effects on AML Cells.Cancers (Basel). 2023 Feb 7;15(4):1039. doi: 10.3390/cancers15041039. Cancers (Basel). 2023. PMID: 36831382 Free PMC article.
-
SLFN11-mediated ribosome biogenesis impairment induces TP53-independent apoptosis.Mol Cell. 2025 Mar 6;85(5):894-912.e10. doi: 10.1016/j.molcel.2025.01.008. Epub 2025 Feb 4. Mol Cell. 2025. PMID: 39909041
-
BRD2 interconnects with BRD3 to facilitate Pol II transcription initiation and elongation to prime promoters for cell differentiation.Cell Mol Life Sci. 2022 Jun 4;79(6):338. doi: 10.1007/s00018-022-04349-4. Cell Mol Life Sci. 2022. PMID: 35665862 Free PMC article.
-
LYAR Promotes Colorectal Cancer Progression by Upregulating FSCN1 Expression and Fatty Acid Metabolism.Oxid Med Cell Longev. 2021 Dec 6;2021:9979707. doi: 10.1155/2021/9979707. eCollection 2021. Oxid Med Cell Longev. 2021. PMID: 35069968 Free PMC article.
References
-
- Ruggero D., Pandolfi P.P.. Does the ribosome translate cancer. Nat. Rev. Cancer. 2003; 3:179–192. - PubMed
-
- White R.J. RNA polymerases I and III, non-coding RNAs and cancer. Trends Genet. 2008; 24:622–629. - PubMed
-
- Boisvert F.M., van Koningsbruggen S., Navascues J., Lamond A.I.. The multifunctional nucleolus. Nat. Rev. Mol. Cell Biol. 2007; 8:574–585. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

