Drosophila melanogaster Mutated in its GBA1b Ortholog Recapitulates Neuronopathic Gaucher Disease
- PMID: 31505865
- PMCID: PMC6780790
- DOI: 10.3390/jcm8091420
Drosophila melanogaster Mutated in its GBA1b Ortholog Recapitulates Neuronopathic Gaucher Disease
Abstract
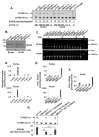
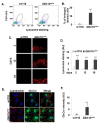
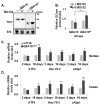
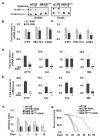
Gaucher disease (GD) results from mutations in the GBA1 gene, which encodes lysosomal glucocerebrosidase (GCase). The large number of mutations known to date in the gene lead to a heterogeneous disorder, which is divided into a non-neuronopathic, type 1 GD, and two neurological, type 2 and type 3, forms. We studied the two fly GBA1 orthologs, GBA1a and GBA1b. Each contains a Minos element insertion, which truncates its coding sequence. In the GBA1am/m flies, which express a mutant protein, missing 33 C-terminal amino acids, there was no decrease in GCase activity or substrate accumulation. However, GBA1bm/m mutant flies presented a significant decrease in GCase activity with concomitant substrate accumulation, which included C14:1 glucosylceramide and C14:0 glucosylsphingosine. GBA1bm/m mutant flies showed activation of the Unfolded Protein Response (UPR) and presented inflammation and neuroinflammation that culminated in development of a neuronopathic disease. Treatment with ambroxol did not rescue GCase activity or reduce substrate accumulation; however, it ameliorated UPR, inflammation and neuroinflammation, and increased life span. Our results highlight the resemblance between the phenotype of the GBA1bm/m mutant fly and neuronopathic GD and underlie its relevance in further GD studies as well as a model to test possible therapeutic modalities.
Keywords: Gaucher disease; GlcCer; GlcSph; glucocerebrosidase; inflammation; unfolded protein response.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Functional Analysis of Human GBA1 Missense Mutations in Drosophila: Insights into Gaucher Disease Pathogenesis and Phenotypic Consequences.Cells. 2024 Sep 27;13(19):1619. doi: 10.3390/cells13191619. Cells. 2024. PMID: 39404383 Free PMC article.
-
The Uncovered Function of the Drosophila GBA1a-Encoded Protein.Cells. 2021 Mar 12;10(3):630. doi: 10.3390/cells10030630. Cells. 2021. PMID: 33809074 Free PMC article.
-
A Drosophila Model of Neuronopathic Gaucher Disease Demonstrates Lysosomal-Autophagic Defects and Altered mTOR Signalling and Is Functionally Rescued by Rapamycin.J Neurosci. 2016 Nov 16;36(46):11654-11670. doi: 10.1523/JNEUROSCI.4527-15.2016. J Neurosci. 2016. PMID: 27852774 Free PMC article.
-
Animal Models for the Study of Gaucher Disease.Int J Mol Sci. 2023 Nov 7;24(22):16035. doi: 10.3390/ijms242216035. Int J Mol Sci. 2023. PMID: 38003227 Free PMC article. Review.
-
Exploring the efficacy and safety of Ambroxol in Gaucher disease: an overview of clinical studies.Front Pharmacol. 2024 Feb 13;15:1335058. doi: 10.3389/fphar.2024.1335058. eCollection 2024. Front Pharmacol. 2024. PMID: 38414738 Free PMC article. Review.
Cited by
-
The Importance of Drosophila melanogaster Research to UnCover Cellular Pathways Underlying Parkinson's Disease.Cells. 2021 Mar 6;10(3):579. doi: 10.3390/cells10030579. Cells. 2021. PMID: 33800736 Free PMC article. Review.
-
Glucocerebrosidase: Functions in and Beyond the Lysosome.J Clin Med. 2020 Mar 9;9(3):736. doi: 10.3390/jcm9030736. J Clin Med. 2020. PMID: 32182893 Free PMC article. Review.
-
Heat shock proteins and small nucleolar RNAs are dysregulated in a Drosophila model for feline hypertrophic cardiomyopathy.G3 (Bethesda). 2021 Jan 18;11(1):jkaa014. doi: 10.1093/g3journal/jkaa014. G3 (Bethesda). 2021. PMID: 33561224 Free PMC article.
-
Lyso-glycosphingolipids: presence and consequences.Essays Biochem. 2020 Sep 23;64(3):565-578. doi: 10.1042/EBC20190090. Essays Biochem. 2020. PMID: 32808655 Free PMC article. Review.
-
'Fly-ing' from rare to common neurodegenerative disease mechanisms.Trends Genet. 2022 Sep;38(9):972-984. doi: 10.1016/j.tig.2022.03.018. Epub 2022 Apr 25. Trends Genet. 2022. PMID: 35484057 Free PMC article. Review.
References
-
- Dekker N., Van Dussen L., Hollak C.E.M., Overkleeft H., Scheij S., Ghauharali K., Van Breemen M.J., Ferraz M.J., Groener J.E.M., Maas M., et al. Elevated plasma glucosylsphingosine in Gaucher disease: Relation to phenotype, storage cell markers, and therapeutic response. Blood. 2011;118:e118–e127. doi: 10.1182/blood-2011-05-352971. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

