Transformation with Artificial Chromosomes in Oxytricha trifallax and Their Applications
- PMID: 31506318
- PMCID: PMC6778790
- DOI: 10.1534/g3.119.400298
Transformation with Artificial Chromosomes in Oxytricha trifallax and Their Applications
Abstract
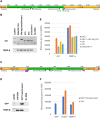
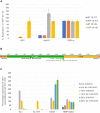
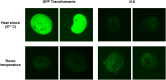
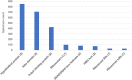
Oxytricha trifallax, like other ciliates, has separate germline and somatic nuclei. The diploid germline genome in the micronucleus is composed of long conventional chromosomes. The macronucleus contains a somatic genome which is naturally fragmented into thousands of kilobase-sized chromosomes. Here, we develop a method to stably incorporate artificial chromosomes into the macronucleus. We report two cases of successful transformation and demonstrate the use of somatic transformation to investigate gene regulation and gene function in Oxytricha We show that the transformed artificial chromosomes are maintained through multiple asexual divisions. Furthermore, they support the transcriptional regulation of the native chromosome from which they were derived and are translated to produce functional proteins. To test if transformed chromosomes are amenable to practical applications, we generated a tagged version of a representative gene (AL1) and used it to co-precipitate associated proteins. This revealed an association with nucleic acid binding proteins, specifically RNA-binding proteins, and RNA immunoprecipitation of AL1 revealed its association with multiple RNAs. The use of artificial chromosomes in Oxytricha enables an array of genetic and molecular biological assays, as well as new avenues of inquiry into the epigenetic programming of macronuclear development and genome rearrangement.
Keywords: Ciliates; Epigenetics; Genome Rearrangement.
Copyright © 2019 Clay et al.
Figures
Similar articles
-
The Oxytricha trifallax macronuclear genome: a complex eukaryotic genome with 16,000 tiny chromosomes.PLoS Biol. 2013;11(1):e1001473. doi: 10.1371/journal.pbio.1001473. Epub 2013 Jan 29. PLoS Biol. 2013. PMID: 23382650 Free PMC article.
-
Programmed Chromosome Deletion in the Ciliate Oxytricha trifallax.G3 (Bethesda). 2019 Oct 7;9(10):3105-3118. doi: 10.1534/g3.118.200930. G3 (Bethesda). 2019. PMID: 31506317 Free PMC article.
-
Mating of the stichotrichous ciliate Oxytricha trifallax induces production of a class of 27 nt small RNAs derived from the parental macronucleus.PLoS One. 2012;7(8):e42371. doi: 10.1371/journal.pone.0042371. Epub 2012 Aug 10. PLoS One. 2012. PMID: 22900016 Free PMC article.
-
Origin, structure and function of millions of chromosomes present in the macronucleus of unicellular eukaryotic ciliate, Oxytricha trifallax: a model organism for transgenerationally programmed genome rearrangements.J Genet. 2015 Jun;94(2):171-6. doi: 10.1007/s12041-015-0504-2. J Genet. 2015. PMID: 26174664 Review. No abstract available.
-
Programmed Genome Rearrangements in the Ciliate Oxytricha.Microbiol Spectr. 2014 Dec;2(6):10.1128/microbiolspec.MDNA3-0025-2014. doi: 10.1128/microbiolspec.MDNA3-0025-2014. Microbiol Spectr. 2014. PMID: 26104449 Free PMC article. Review.
Cited by
-
Application of RNA interference and protein localization to investigate housekeeping and developmentally regulated genes in the emerging model protozoan Paramecium caudatum.Commun Biol. 2024 Feb 19;7(1):204. doi: 10.1038/s42003-024-05906-2. Commun Biol. 2024. PMID: 38374195 Free PMC article.
-
Relaxed DNA substrate specificity of transposases involved in programmed genome rearrangement.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf577. doi: 10.1093/nar/gkaf577. Nucleic Acids Res. 2025. PMID: 40637233 Free PMC article.
-
Relaxed DNA substrate specificity of transposases involved in programmed genome rearrangement.bioRxiv [Preprint]. 2025 Mar 18:2025.03.17.643836. doi: 10.1101/2025.03.17.643836. bioRxiv. 2025. Update in: Nucleic Acids Res. 2025 Jul 8;53(13):gkaf577. doi: 10.1093/nar/gkaf577. PMID: 40166331 Free PMC article. Updated. Preprint.
References
-
- Bulic A., Postberg J., Fischer A., Jönsson F., Reuter G. et al. , 2013. A permissive chromatin structure is adopted prior to site-specific DNA demethylation of developmentally expressed genes involved in macronuclear differentiation. Epigenetics Chromatin 6: 5 10.1186/1756-8935-6-5 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
