Pharmaco-Genetic Screen To Uncover Actin Regulators Targeted by Prostaglandins During Drosophila Oogenesis
- PMID: 31506320
- PMCID: PMC6829128
- DOI: 10.1534/g3.119.400704
Pharmaco-Genetic Screen To Uncover Actin Regulators Targeted by Prostaglandins During Drosophila Oogenesis
Abstract
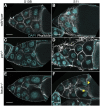
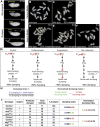
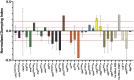
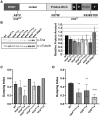
Prostaglandins (PGs) are lipid signaling molecules with numerous physiologic functions, including pain/inflammation, fertility, and cancer. PGs are produced downstream of cyclooxygenase (COX) enzymes, the targets of non-steroidal anti-inflammatory drugs (NSAIDs). In numerous systems, PGs regulate actin cytoskeletal remodeling, however, their mechanisms of action remain largely unknown. To address this deficiency, we undertook a pharmaco-genetic interaction screen during late-stage Drosophila oogenesis. Drosophila oogenesis is as an established model for studying both actin dynamics and PGs. Indeed, during Stage 10B, cage-like arrays of actin bundles surround each nurse cell nucleus, and during Stage 11, the cortical actin contracts, squeezing the cytoplasmic contents into the oocyte. Both of these cytoskeletal properties are required for follicle development and fertility, and are regulated by PGs. Here we describe a pharmaco-genetic interaction screen that takes advantage of the fact that Stage 10B follicles will mature in culture and COX inhibitors, such as aspirin, block this in vitro follicle maturation. In the screen, aspirin was used at a concentration that blocks 50% of the wild-type follicles from maturing in culture. By combining this aspirin treatment with heterozygosity for mutations in actin regulators, we quantitatively identified enhancers and suppressors of COX inhibition. Here we present the screen results and initial follow-up studies on three strong enhancers - Enabled, Capping protein, and non-muscle Myosin II Regulatory Light Chain. Overall, these studies provide new insight into how PGs regulate both actin bundle formation and cellular contraction, properties that are not only essential for development, but are misregulated in disease.
Keywords: Capping protein; Drosophila melanogaster; Enabled; non-muscle myosin II regulatory light chain; prostaglandins.
Copyright © 2019 Spracklen et al.
Figures
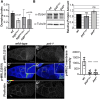
References
-
- Abramoff M. D., Magalhaes P. J., and Ram S. J., 2004. Image Processing with ImageJ. Biophoton. Int. 11: 36–42.
-
- Ahern-Djamali S. M., Comer A. R., Bachmann C., Kastenmeier A. S., Reddy S. K. et al. , 1998. Mutations in Drosophila enabled and rescue by human vasodilator-stimulated phosphoprotein (VASP) indicate important functional roles for Ena/VASP homology domain 1 (EVH1) and EVH2 domains. Mol. Biol. Cell 9: 2157–2171. 10.1091/mbc.9.8.2157 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
