Transcript Analysis of Zebrafish GLUT3 Genes, slc2a3a and slc2a3b, Define Overlapping as Well as Distinct Expression Domains in the Zebrafish (Danio rerio) Central Nervous System
- PMID: 31507372
- PMCID: PMC6718831
- DOI: 10.3389/fnmol.2019.00199
Transcript Analysis of Zebrafish GLUT3 Genes, slc2a3a and slc2a3b, Define Overlapping as Well as Distinct Expression Domains in the Zebrafish (Danio rerio) Central Nervous System
Abstract
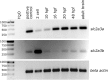
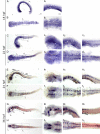
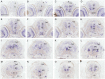
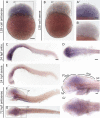
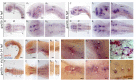
The transport of glucose across the cell plasma membrane is vital to most mammalian cells. The glucose transporter (GLUT; also called SLC2A) family of transmembrane solute carriers is responsible for this function in vivo. GLUT proteins encompass 14 different isoforms in humans with different cell type-specific expression patterns and activities. Central to glucose utilization and delivery in the brain is the neuronally expressed GLUT3. Recent research has shown an involvement of GLUT3 genetic variation or altered expression in several different brain disorders, including Huntington's and Alzheimer's diseases. Furthermore, GLUT3 was identified as a potential risk gene for multiple psychiatric disorders. To study the role of GLUT3 in brain function and disease a more detailed knowledge of its expression in model organisms is needed. Zebrafish (Danio rerio) has in recent years gained popularity as a model organism for brain research and is now well-established for modeling psychiatric disorders. Here, we have analyzed the sequence of GLUT3 orthologs and identified two paralogous genes in the zebrafish, slc2a3a and slc2a3b. Interestingly, the Glut3b protein sequence contains a unique stretch of amino acids, which may be important for functional regulation. The slc2a3a transcript is detectable in the central nervous system including distinct cellular populations in telencephalon, diencephalon, mesencephalon and rhombencephalon at embryonic and larval stages. Conversely, the slc2a3b transcript shows a rather diffuse expression pattern at different embryonic stages and brain regions. Expression of slc2a3a is maintained in the adult brain and is found in the telencephalon, diencephalon, mesencephalon, cerebellum and medulla oblongata. The slc2a3b transcripts are present in overlapping as well as distinct regions compared to slc2a3a. Double in situ hybridizations were used to demonstrate that slc2a3a is expressed by some GABAergic neurons at embryonic stages. This detailed description of zebrafish slc2a3a and slc2a3b expression at developmental and adult stages paves the way for further investigations of normal GLUT3 function and its role in brain disorders.
Keywords: GABA; GAD1; brain development; brain disorders; glucose transporter; nervous system; psychiatric disorders.
Figures




Similar articles
-
Distribution of transcripts of the GFOD gene family members gfod1 and gfod2 in the zebrafish central nervous system.Gene Expr Patterns. 2020 Jun;36:119111. doi: 10.1016/j.gep.2020.119111. Epub 2020 Mar 18. Gene Expr Patterns. 2020. PMID: 32197942
-
GLUT3 gene expression is critical for embryonic growth, brain development and survival.Mol Genet Metab. 2014 Apr;111(4):477-83. doi: 10.1016/j.ymgme.2014.01.013. Epub 2014 Jan 31. Mol Genet Metab. 2014. PMID: 24529979 Free PMC article.
-
Glial cell line-derived neurotrophic factor expression in the brain of adult zebrafish (Danio rerio).Histol Histopathol. 2008 Mar;23(3):251-61. doi: 10.14670/HH-23.251. Histol Histopathol. 2008. PMID: 18072082
-
Current State of Modeling Human Psychiatric Disorders Using Zebrafish.Int J Mol Sci. 2023 Feb 6;24(4):3187. doi: 10.3390/ijms24043187. Int J Mol Sci. 2023. PMID: 36834599 Free PMC article. Review.
-
Can we gain translational insights into the functional roles of cerebral cortex from acortical rodent and naturally acortical zebrafish models?Prog Neuropsychopharmacol Biol Psychiatry. 2024 Jun 8;132:110964. doi: 10.1016/j.pnpbp.2024.110964. Epub 2024 Feb 13. Prog Neuropsychopharmacol Biol Psychiatry. 2024. PMID: 38354895 Review.
Cited by
-
Loss-of-Function Models of the Metabotropic Glutamate Receptor Genes Grm8a and Grm8b Display Distinct Behavioral Phenotypes in Zebrafish Larvae (Danio rerio).Front Mol Neurosci. 2022 Jun 13;15:901309. doi: 10.3389/fnmol.2022.901309. eCollection 2022. Front Mol Neurosci. 2022. PMID: 35769333 Free PMC article.
-
Zebrafish, Medaka and Turquoise Killifish for Understanding Human Neurodegenerative/Neurodevelopmental Disorders.Int J Mol Sci. 2022 Jan 26;23(3):1399. doi: 10.3390/ijms23031399. Int J Mol Sci. 2022. PMID: 35163337 Free PMC article. Review.
-
Oxidative Stress, Inflammation and Altered Glucose Metabolism Contribute to the Retinal Phenotype in the Choroideremia Zebrafish.Antioxidants (Basel). 2024 Dec 23;13(12):1587. doi: 10.3390/antiox13121587. Antioxidants (Basel). 2024. PMID: 39765914 Free PMC article.
References
-
- Amir Shaghaghi M., Zhouyao H., Tu H., El-Gabalawy H., Crow G. H., Levine M., et al. . (2017). The SLC2A14 gene, encoding the novel glucose/dehydroascorbate transporter GLUT14, is associated with inflammatory bowel disease. Am. J. Clin. Nutr. 106, 1508–1513. 10.3945/ajcn.116.147603 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

