Origins and Evolution of the α-L-Fucosidases: From Bacteria to Metazoans
- PMID: 31507539
- PMCID: PMC6718869
- DOI: 10.3389/fmicb.2019.01756
Origins and Evolution of the α-L-Fucosidases: From Bacteria to Metazoans
Abstract
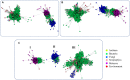
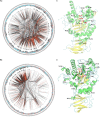
α-L-fucosidases (EC 3.2.1.51, FUC), belonging to the glycoside hydrolase family 29 (GH29), play important roles in several biological processes and are markers used for detecting hepatocellular carcinoma. In this study, a protein sequence similarity network (SSN) was generated and a subsequent evolutionary analysis was performed to understand the enzymes comprehensively. The SSN indicated that the proteins in the FUC family are mainly present in bacteria, fungi, metazoans, plants, as well as in archaea, but less abundantly. The sequences in bacteria were found to be more diverse than those in other taxonomic groups. The SSN and a phylogenetic tree both supported that the proteins in the FUC family can be classified into 3 subfamilies. FUCs in each subfamily are under the pressure of negative selection. The enzymes from metazoans, fungi, and plants separated into the three subfamilies and shared high similarity with the bacterial homologs. The multiple sequence alignment results indicated that the amino acid residues for binding α-L-fucosidase and catalysis are highly conserved in the 3 subfamilies; however, the evolutionary patterns were different, based on the coevolution analysis in the subfamily of metazoans and bacteria. Finally, gene duplication plays an important role for α-L-fucosidase evolution, not only in metazoans, but also in bacteria and fungi.
Keywords: bacteria; evolution; metazoa; sequence similarity network; α-L-fucosidase.
Figures



Similar articles
-
Substrate specificity and transglycosylation capacity of α-L-fucosidases across GH29 assessed by bioinformatics-assisted selection of functional diversity.Glycobiology. 2023 Jun 3;33(5):396-410. doi: 10.1093/glycob/cwad029. Glycobiology. 2023. PMID: 37014745
-
Fucoidan-active α-L-fucosidases of the GH29 and GH95 families from a fucoidan degrading cluster of the marine bacterium Wenyingzhuangia fucanilytica.Arch Biochem Biophys. 2022 Oct 15;728:109373. doi: 10.1016/j.abb.2022.109373. Epub 2022 Aug 5. Arch Biochem Biophys. 2022. PMID: 35940339
-
Characterization of five marine family 29 glycoside hydrolases reveals an α-L-fucosidase targeting specifically Fuc(α1,4)GlcNAc.Glycobiology. 2022 May 23;32(6):529-539. doi: 10.1093/glycob/cwab132. Glycobiology. 2022. PMID: 35137077
-
Exploring the sequence-function space of microbial fucosidases.Commun Chem. 2024 Jun 18;7(1):137. doi: 10.1038/s42004-024-01212-4. Commun Chem. 2024. PMID: 38890439 Free PMC article.
-
The alpha-L-fucosidase from Sulfolobus solfataricus.Extremophiles. 2008 Jan;12(1):61-8. doi: 10.1007/s00792-007-0105-y. Epub 2007 Aug 9. Extremophiles. 2008. PMID: 17687508 Review.
Cited by
-
Architecture Insight of Bifidobacterial α-L-Fucosidases.Int J Mol Sci. 2021 Aug 6;22(16):8462. doi: 10.3390/ijms22168462. Int J Mol Sci. 2021. PMID: 34445166 Free PMC article.
-
Comparative genomics of Clostridium butyricum reveals a conserved genome architecture and novel virulence-related gene clusters.Microb Genom. 2025 Aug;11(8):001477. doi: 10.1099/mgen.0.001477. Microb Genom. 2025. PMID: 40794526 Free PMC article.
-
The dual role of fucosidases: tool or target.Biologia (Bratisl). 2023 Feb 28:1-16. doi: 10.1007/s11756-023-01351-4. Online ahead of print. Biologia (Bratisl). 2023. PMID: 37363646 Free PMC article. Review.
-
Development of an Autochthonous Microbial Consortium for Enhanced Bioremediation of PAH-Contaminated Soil.Int J Mol Sci. 2021 Dec 15;22(24):13469. doi: 10.3390/ijms222413469. Int J Mol Sci. 2021. PMID: 34948267 Free PMC article.
-
Synthesis of fucosyllactose using α-L-fucosidases GH29 from infant gut microbial metagenome.Appl Microbiol Biotechnol. 2024 May 21;108(1):338. doi: 10.1007/s00253-024-13178-3. Appl Microbiol Biotechnol. 2024. PMID: 38771321 Free PMC article.
References
LinkOut - more resources
Full Text Sources

