Shaping Ethylene Response: The Role of EIN3/EIL1 Transcription Factors
- PMID: 31507622
- PMCID: PMC6718143
- DOI: 10.3389/fpls.2019.01030
Shaping Ethylene Response: The Role of EIN3/EIL1 Transcription Factors
Abstract
EIN3/EIL1 transcription factors are the key regulators of ethylene signaling that sustain a variety of plant responses to ethylene. Since ethylene regulates multiple aspects of plant development and stress responses, its signaling outcome needs proper modulation depending on the spatiotemporal and environmental conditions. In this review, we summarize recent advances on the molecular mechanisms that underlie EIN3/EIL1-directed ethylene signaling in Arabidopsis. We focus on the role of EIN3/EIL1 in tuning transcriptional regulation of ethylene response in time and space. Besides, we consider the role of EIN3/EIL1-independent regulation of ethylene signaling.
Keywords: ETHYLENE-INSENSITIVE3; ETHYLENE-INSENSITIVE3-LIKE; cross-talk; epigenetic regulation; protein–protein interactions.
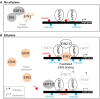
Figures


Similar articles
-
Ethylene-induced stabilization of ETHYLENE INSENSITIVE3 and EIN3-LIKE1 is mediated by proteasomal degradation of EIN3 binding F-box 1 and 2 that requires EIN2 in Arabidopsis.Plant Cell. 2010 Jul;22(7):2384-401. doi: 10.1105/tpc.110.076588. Epub 2010 Jul 20. Plant Cell. 2010. PMID: 20647342 Free PMC article.
-
Integrated Regulation of Apical Hook Development by Transcriptional Coupling of EIN3/EIL1 and PIFs in Arabidopsis.Plant Cell. 2018 Sep;30(9):1971-1988. doi: 10.1105/tpc.18.00018. Epub 2018 Aug 13. Plant Cell. 2018. PMID: 30104405 Free PMC article.
-
The Arabidopsis EIN3 binding F-Box proteins EBF1 and EBF2 have distinct but overlapping roles in ethylene signaling.Plant Cell. 2007 Feb;19(2):509-23. doi: 10.1105/tpc.106.048140. Epub 2007 Feb 16. Plant Cell. 2007. PMID: 17307926 Free PMC article.
-
Paradigms and paradox in the ethylene signaling pathway and interaction network.Mol Plant. 2011 Jul;4(4):626-34. doi: 10.1093/mp/ssr042. Epub 2011 Jun 20. Mol Plant. 2011. PMID: 21690206 Review.
-
The ethylene signaling pathway: new insights.Curr Opin Plant Biol. 2004 Feb;7(1):40-9. doi: 10.1016/j.pbi.2003.11.011. Curr Opin Plant Biol. 2004. PMID: 14732440 Review.
Cited by
-
Comparison of the pathway structures influencing the temporal response of salicylate and jasmonate defence hormones in Arabidopsis thaliana.Front Plant Sci. 2022 Sep 9;13:952301. doi: 10.3389/fpls.2022.952301. eCollection 2022. Front Plant Sci. 2022. PMID: 36160984 Free PMC article. Review.
-
Essential Roles of the Linker Sequence Between Tetratricopeptide Repeat Motifs of Ethylene Overproduction 1 in Ethylene Biosynthesis.Front Plant Sci. 2021 Apr 15;12:657300. doi: 10.3389/fpls.2021.657300. eCollection 2021. Front Plant Sci. 2021. PMID: 33936142 Free PMC article.
-
A guide to plant TPX2-like and WAVE-DAMPENED2-like proteins.J Exp Bot. 2021 Feb 24;72(4):1034-1045. doi: 10.1093/jxb/eraa513. J Exp Bot. 2021. PMID: 33130902 Free PMC article.
-
Chloroplast Vesiculation and Induced Chloroplast Vesiculation and Senescence-Associated Gene 12 Expression During Tomato Flower Pedicel Abscission.Plant Direct. 2025 Jan 8;9(1):e70035. doi: 10.1002/pld3.70035. eCollection 2025 Jan. Plant Direct. 2025. PMID: 39790709 Free PMC article.
-
Limited conservation in cross-species comparison of GLK transcription factor binding suggested wide-spread cistrome divergence.Nat Commun. 2022 Dec 9;13(1):7632. doi: 10.1038/s41467-022-35438-4. Nat Commun. 2022. PMID: 36494366 Free PMC article.
References
-
- Abeles F. B., Morgan P. W., Saltveit M. E., Jr. (2012). Ethylene in plant biology. Academic press; San Diego, CA.
-
- Alonso J. M., Stepanova A. N., Solano R., Wisman E., Ferrari S., Ausubel F. M., et al. (2003). Five components of the ethylene-response pathway identified in a screen for weak ethylene-insensitive mutants in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 100, 2992–1997. 10.1073/pnas.0438070100 - DOI - PMC - PubMed
-
- An F., Zhao Q., Ji Y., Li W., Jiang Z., Yu X., et al. (2010). Ethylene-induced stabilization of ETHYLENE INSENSITIVE3 and EIN3-LIKE1 is mediated by proteasomal degradation of EIN3 binding F-box 1 and 2 that requires EIN2 in Arabidopsis. Plant Cell 22, 2384–2401. 10.1105/tpc.110.076588 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

