ER stress-linked autophagy stabilizes apoptosis effector PERP and triggers its co-localization with SERCA2b at ER-plasma membrane junctions
- PMID: 31508245
- PMCID: PMC6718399
- DOI: 10.1038/s41420-019-0212-4
ER stress-linked autophagy stabilizes apoptosis effector PERP and triggers its co-localization with SERCA2b at ER-plasma membrane junctions
Abstract
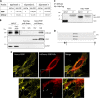
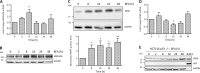
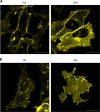
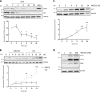
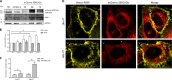
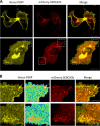
Specific molecular interactions that underpin the switch between ER stress-triggered autophagy-mediated cellular repair and cellular death by apoptosis are not characterized. This study reports the unexpected interaction elicited by ER stress between the plasma membrane (PM)-localized apoptosis effector PERP and the ER Ca2+ pump SERCA2b. We show that the p53 effector PERP, which specifically induces apoptosis when expressed above a threshold level, has a heterogeneous distribution across the PM of un-stressed cells and is actively turned over by the lysosome. PERP is upregulated following sustained starvation-induced autophagy, which precedes the onset of apoptosis indicating that PERP protein levels are controlled by a lysosomal pathway that is sensitive to cellular physiological state. Furthermore, ER stress stabilizes PERP at the PM and induces its increasing co-localization with SERCA2b at ER-PM junctions. The findings highlight a novel crosstalk between pro-survival autophagy and pro-death apoptosis pathways and identify, for the first time, accumulation of an apoptosis effector to ER-PM junctions in response to ER stress.
Keywords: Apoptosis; Lysosomes; Macroautophagy.
Conflict of interest statement
Conflict of interestThe authors declare that they have no conflict of interest.
Figures

Similar articles
-
PERP-ing into diverse mechanisms of cancer pathogenesis: Regulation and role of the p53/p63 effector PERP.Biochim Biophys Acta Rev Cancer. 2020 Aug;1874(1):188393. doi: 10.1016/j.bbcan.2020.188393. Epub 2020 Jul 15. Biochim Biophys Acta Rev Cancer. 2020. PMID: 32679166 Review.
-
PERP, an apoptosis-associated target of p53, is a novel member of the PMP-22/gas3 family.Genes Dev. 2000 Mar 15;14(6):704-18. Genes Dev. 2000. PMID: 10733530 Free PMC article.
-
PERP expression stabilizes active p53 via modulation of p53-MDM2 interaction in uveal melanoma cells.Cell Death Dis. 2011 Mar 31;2(3):e136. doi: 10.1038/cddis.2011.19. Cell Death Dis. 2011. PMID: 21451571 Free PMC article.
-
P53 apoptosis mediator PERP: localization, function and caspase activation in uveal melanoma.J Cell Mol Med. 2009 Aug;13(8B):1995-2007. doi: 10.1111/j.1582-4934.2008.00590.x. J Cell Mol Med. 2009. PMID: 19040420 Free PMC article.
-
Regulation and Role of Store-Operated Ca2+ Entry in Cellular Proliferation.In: Kozak JA, Putney JW Jr, editors. Calcium Entry Channels in Non-Excitable Cells. Boca Raton (FL): CRC Press/Taylor & Francis; 2018. Chapter 12. In: Kozak JA, Putney JW Jr, editors. Calcium Entry Channels in Non-Excitable Cells. Boca Raton (FL): CRC Press/Taylor & Francis; 2018. Chapter 12. PMID: 30299656 Free Books & Documents. Review.
Cited by
-
Endoplasmic Reticulum Stress in Bronchopulmonary Dysplasia: Contributor or Consequence?Cells. 2024 Oct 26;13(21):1774. doi: 10.3390/cells13211774. Cells. 2024. PMID: 39513884 Free PMC article. Review.
-
Water-based propolis enhances 5-fluorouracil drug efficiency in gastric and colorectal cancer cells through cell stress response, anti-migratory, and apoptotic effects regardless of p53 status.Med Oncol. 2025 Aug 26;42(10):449. doi: 10.1007/s12032-025-03023-6. Med Oncol. 2025. PMID: 40858788
-
Upregulation of METTL14 mediates the elevation of PERP mRNA N6 adenosine methylation promoting the growth and metastasis of pancreatic cancer.Mol Cancer. 2020 Aug 25;19(1):130. doi: 10.1186/s12943-020-01249-8. Mol Cancer. 2020. PMID: 32843065 Free PMC article.
-
Perspectives on Organelle Interaction, Protein Dysregulation, and Cancer Disease.Front Cell Dev Biol. 2021 Feb 25;9:613336. doi: 10.3389/fcell.2021.613336. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 33718356 Free PMC article. Review.
-
P53 and BCL-2 family proteins PUMA and NOXA define competitive fitness in pluripotent cell competition.PLoS Genet. 2024 Mar 15;20(3):e1011193. doi: 10.1371/journal.pgen.1011193. eCollection 2024 Mar. PLoS Genet. 2024. PMID: 38489392 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

