Structural and functional analysis of the Hsp70/Hsp40 chaperone system
- PMID: 31509306
- PMCID: PMC6954727
- DOI: 10.1002/pro.3725
Structural and functional analysis of the Hsp70/Hsp40 chaperone system
Abstract
As one of the most abundant and highly conserved molecular chaperones, the 70-kDa heat shock proteins (Hsp70s) play a key role in maintaining cellular protein homeostasis (proteostasis), one of the most fundamental tasks for every living organism. In this role, Hsp70s are inextricably linked to many human diseases, most notably cancers and neurodegenerative diseases, and are increasingly recognized as important drug targets for developing novel therapeutics for these diseases. Hsp40s are a class of essential and universal partners for Hsp70s in almost all aspects of proteostasis. Thus, Hsp70s and Hsp40s together constitute one of the most important chaperone systems across all kingdoms of life. In recent years, we have witnessed significant progress in understanding the molecular mechanism of this chaperone system through structural and functional analysis. This review will focus on this recent progress, mainly from a structural perspective.
Keywords: Hsp40; Hsp70; molecular chaperone; neurodegenerative diseases; protein folding; proteostasis.
© 2019 The Protein Society.
Conflict of interest statement
The authors declare no potential conflict of interest.
Figures

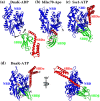

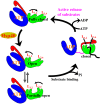
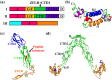
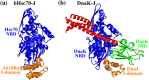
References
-
- Hartl FU, Hayer‐Hartl M. Converging concepts of protein folding in vitro and in vivo. Nat Struct Mol Biol. 2009;16:574–581. - PubMed
-
- Bukau B, Weissman J, Horwich A. Molecular chaperones and protein quality control. Cell. 2006;125:443–451. - PubMed
-
- Bukau B, Deuerling E, Pfund C, Craig EA. Getting newly synthesized proteins into shape. Cell. 2000;101:119–122. - PubMed
-
- Bukau B, Horwich AL. The Hsp70 and Hsp60 chaperone machines. Cell. 1998;92:351–366. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

