An Adaptive Real-Time 3D Single Particle Tracking Method for Monitoring Viral First Contacts
- PMID: 31529595
- PMCID: PMC6824287
- DOI: 10.1002/smll.201903039
An Adaptive Real-Time 3D Single Particle Tracking Method for Monitoring Viral First Contacts
Abstract
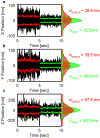
Here, an adaptive real-time 3D single particle tracking method is proposed, which is capable of capturing heterogeneous dynamics. Using a real-time measurement of a rapidly diffusing particle's positional variance, the 3D precision adaptive real-time tracking (3D-PART) microscope adjusts active-feedback parameters to trade tracking speed for precision on demand. This technique is demonstrated first on immobilized fluorescent nanoparticles, with a greater than twofold increase in the lateral localization precision (≈25 to ≈11 nm at 1 ms sampling) as well as a smaller increase in the axial localization precision (≈ 68 to ≈45 nm). 3D-PART also shows a marked increase in the precision when tracking freely diffusing particles, with lateral precision increasing from ≈100 to ≈70 nm for particles diffusing at 4 µm2 s-1 , although with a sacrifice in the axial precision (≈250 to ≈350 nm). This adaptive microscope is then applied to monitoring the viral first contacts of virus-like particles to the surface of live cells, allowing direct and continuous measurement of the viral particle at initial contact with the cell surface.
Keywords: 3D microscopy; adaptive methods; real-time single particle tracking; virus tracking.
© 2019 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures
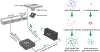





Similar articles
-
Real-Time 3D Single Particle Tracking: Towards Active Feedback Single Molecule Spectroscopy in Live Cells.Molecules. 2019 Aug 2;24(15):2826. doi: 10.3390/molecules24152826. Molecules. 2019. PMID: 31382495 Free PMC article. Review.
-
Real-time parallel 3D multiple particle tracking with single molecule centrifugal force microscopy.J Microsc. 2019 Mar;273(3):178-188. doi: 10.1111/jmi.12773. Epub 2018 Nov 29. J Microsc. 2019. PMID: 30489640
-
Robust real-time 3D single-particle tracking using a dynamically moving laser spot.Opt Lett. 2017 Jun 15;42(12):2390-2393. doi: 10.1364/OL.42.002390. Opt Lett. 2017. PMID: 28614318
-
Three-Dimensional Tracking of Tethered Particles for Probing Nanometer-Scale Single-Molecule Dynamics Using a Plasmonic Microscope.ACS Sens. 2021 Nov 26;6(11):4234-4243. doi: 10.1021/acssensors.1c01927. Epub 2021 Nov 17. ACS Sens. 2021. PMID: 34786931 Free PMC article.
-
Three-Dimensional Localization of Single Molecules for Super-Resolution Imaging and Single-Particle Tracking.Chem Rev. 2017 Jun 14;117(11):7244-7275. doi: 10.1021/acs.chemrev.6b00629. Epub 2017 Feb 2. Chem Rev. 2017. PMID: 28151646 Free PMC article. Review.
Cited by
-
Recent Advances in Single-Molecule Tracking and Imaging Techniques.Annu Rev Anal Chem (Palo Alto Calif). 2023 Jun 14;16(1):253-284. doi: 10.1146/annurev-anchem-091922-073057. Annu Rev Anal Chem (Palo Alto Calif). 2023. PMID: 37314878 Free PMC article. Review.
-
Particle-by-Particle In Situ Characterization of the Protein Corona via Real-Time 3D Single-Particle-Tracking Spectroscopy*.Angew Chem Int Ed Engl. 2021 Oct 4;60(41):22359-22367. doi: 10.1002/anie.202105741. Epub 2021 Aug 1. Angew Chem Int Ed Engl. 2021. PMID: 34015174 Free PMC article.
-
Capturing the start point of the virus-cell interaction with high-speed 3D single-virus tracking.Nat Methods. 2022 Dec;19(12):1642-1652. doi: 10.1038/s41592-022-01672-3. Epub 2022 Nov 10. Nat Methods. 2022. PMID: 36357694 Free PMC article.
-
Combined online Bayesian and windowed estimation of background and signal localization facilitates active-feedback particle tracking in complex environments.J Chem Phys. 2022 Nov 14;157(18):184108. doi: 10.1063/5.0118317. J Chem Phys. 2022. PMID: 36379789 Free PMC article.
-
Real-Time Feedback-Driven Single-Particle Tracking: A Survey and Perspective.Small. 2022 Jul;18(29):e2107024. doi: 10.1002/smll.202107024. Epub 2022 Jun 27. Small. 2022. PMID: 35758534 Free PMC article. Review.
References
-
- Cang H; Xu CS; Montiel D; Yang H, Opt Lett 2007, 32, 2729; - PubMed
- Cang H; Wong CM; Xu CS; Rizvi AH; Yang H, Applied Physics Letters 2006, 88, 223901;
- Lessard GA; Goodwin PM; Werner JH, Applied Physics Letters 2007, 91, 224106;
- Katayama Y; Burkacky O; Meyer M; Brauchle C; Gratton E; Lamb DC, Chemphyschem 2009, 10, 2458; - PMC - PubMed
- Juette MF; Bewersdorf J, Nano Lett 2010, 10, 4657; - PubMed
- McHale K; Berglund AJ; Mabuchi H, Nano Lett 2007, 7, 3535; - PubMed
- Levi V; Ruan Q; Gratton E, Biophys J 2005, 88, 2919; - PMC - PubMed
- Perillo EP; Liu YL; Huynh K; Liu C; Chou CK; Hung MC; Yeh HC; Dunn AK, Nat Commun 2015, 6, 7874; - PMC - PubMed
- Hou; Lang; Welsher, Opt Lett 2017, 42, 2390–2393; - PubMed
- Welsher K; Yang H, Nat Nanotechnol 2014, 9, 198. - PubMed
-
- Mermillod-Blondin A; McLeod E; Arnold CB, Opt Lett 2008, 33, 2146. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

