A real-time Monte Carlo tool for individualized dose estimations in clinical CT
- PMID: 31539892
- PMCID: PMC7050822
- DOI: 10.1088/1361-6560/ab467f
A real-time Monte Carlo tool for individualized dose estimations in clinical CT
Abstract
The increasing awareness of the adverse effects associated with radiation exposure in computed tomography (CT) has necessesitated the quantification of dose delivered to patients for better risk assessment in the clinic. The current methods for dose quantification used in the clinic are approximations, lacking realistic models for the irradiation conditions utilized in the scan and the anatomy of the patient being imaged, which limits their relevance for a particular patient. The established gold-standard technique for individualized dose quantification uses Monte Carlo (MC) simulations to obtain patient-specific estimates of organ dose in anatomically realistic computational phantoms to provide patient-specific estimates of organ dose. Although accurate, MC simulations are computationally expensive, which limits their utility for time-constrained applications in the clinic. To overcome these shortcomings, a real-time GPU-based MC tool based on FDA's MC-GPU framework was developed for patient and scanner-specific dosimetry in the clinic. The tool was validated against (1) AAPM's TG-195 reference datasets and (2) physical measurements of dose acquired using TLD chips in adult and pediatric anthropomorphic phantoms. To demonstrate its utility towards providing individualized dose estimates, it was integrated with an automatic segmentation method for generating patient-specific models, which were then used to estimate patient- and scanner-specific organ doses for a select population of 50 adult patients using a clinically relevant CT protocol. The organ dose estimates were compared to corresponding dose estimates from a previously validated MC method based on Penelope. The dose estimates from our MC tool agreed within 5% for all organs (except thyroid) tabulated by TG-195 and within 10% for all TLD locations in the adult and pediactric phantoms, across all validation cases. Compared against Penelope, the organ dose estimates agreed within 3% on average for all organs in the patient population study. The average run duration for each patient was estimated at 23.79 s, representing a significant speedup (~700×) over existing non-parallelized MC methods. The accuracy of dose estimates combined with a significant improvement in execution times suggests a feasible solution utilizing the proposed MC tool for real-time individualized dosimetry in the clinic.
Figures


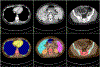
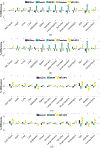

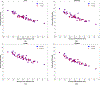
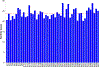
References
-
- Badal A and Badano A 2009. Accelerating Monte Carlo simulations of photon transport in a voxelized geometry using a massively parallel graphics processing unit Med. Phys 36 4878–80 - PubMed
-
- Brink J A and Morin R L 2012. Size-specific dose estimation for CT: how should it be used and what does it mean? Radiology 265 666–8 - PubMed
-
- Chen W, Kolditz D, Beister M, Bohle R and Kalender W A 2012. Fast on-site Monte Carlo tool for dose calculations in CT applications Med. Phys 39 2985–96 - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous
