Systematic Identification of Host Cell Regulators of Legionella pneumophila Pathogenesis Using a Genome-wide CRISPR Screen
- PMID: 31540829
- PMCID: PMC6800164
- DOI: 10.1016/j.chom.2019.08.017
Systematic Identification of Host Cell Regulators of Legionella pneumophila Pathogenesis Using a Genome-wide CRISPR Screen
Abstract
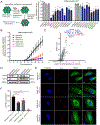
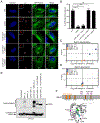
During infection, Legionella pneumophila translocates over 300 effector proteins into the host cytosol, allowing the pathogen to establish an endoplasmic reticulum (ER)-like Legionella-containing vacuole (LCV) that supports bacterial replication. Here, we perform a genome-wide CRISPR-Cas9 screen and secondary targeted screens in U937 human monocyte/macrophage-like cells to systematically identify host factors that regulate killing by L. pneumophila. The screens reveal known host factors hijacked by L. pneumophila, as well as genes spanning diverse trafficking and signaling pathways previously not linked to L. pneumophila pathogenesis. We further characterize C1orf43 and KIAA1109 as regulators of phagocytosis and show that RAB10 and its chaperone RABIF are required for optimal L. pneumophila replication and ER recruitment to the LCV. Finally, we show that Rab10 protein is recruited to the LCV and ubiquitinated by the effectors SidC/SdcA. Collectively, our results provide a wealth of previously undescribed insights into L. pneumophila pathogenesis and mammalian cell function.
Keywords: C1ORF43; CRISPR screen; Legionella pneumophila; RAB10; host-pathogen interaction; intracellular bacteria; pathogen; phagocytosis.
Copyright © 2019 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests
The authors declare no competing interests.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

