The Discovery of RNA Aptamers that Selectively Bind Glioblastoma Stem Cells
- PMID: 31541799
- PMCID: PMC6796606
- DOI: 10.1016/j.omtn.2019.08.015
The Discovery of RNA Aptamers that Selectively Bind Glioblastoma Stem Cells
Abstract
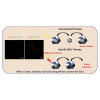
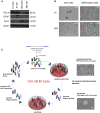
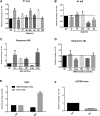
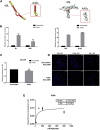
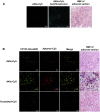
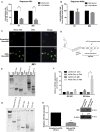
Glioblastoma (GBM) is the most aggressive primary brain tumor in adults. Despite progress in surgical and medical neuro-oncology, prognosis for GBM patients remains dismal, with a median survival of only 14-15 months. The modest benefit of conventional therapies is due to the presence of GBM stem cells (GSCs) that cause tumor relapse and chemoresistance and, therefore, that play a key role in GBM aggressiveness and recurrence. So far, strategies to identify and target GSCs have been unsuccessful. Thus, the development of an approach for GSC detection and targeting would be fundamental for improving the survival of GBM patients. Here, using the cell-systematic evolution of ligand by exponential (SELEX) methodology on human primary GSCs, we generated and characterized RNA aptamers that selectively bind GSCs versus undifferentiated GBM cells. We found that the shortened version of the aptamer 40L, which we have called A40s, costained with CD133-labeled cells in human GBM tissue, suggestive of an ability to specifically recognize GSCs in fixed human tissues. Of note, both 40L and A40s were rapidly internalized by cells, allowing for the delivery of the microRNA miR-34c and the anti-microRNA anti-miR-10b, demonstrating that these aptamers can serve as selective vehicles for therapeutics. In conclusion, the aptamers 40L and A40s can selectively target GSCs. Given the crucial role of GSCs in GBM recurrence and therapy resistance, these aptamers represent innovative drug delivery candidates with a great potential in the treatment of GBM.
Keywords: aptamer; cancer stem cell; delivery; glioblastoma; microRNA.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures

References
-
- Louis D.N., Perry A., Reifenberger G., von Deimling A., Figarella-Branger D., Cavenee W.K., Ohgaki H., Wiestler O.D., Kleihues P., Ellison D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol. 2016;131:803–820. - PubMed
-
- Sant M., Minicozzi P., Lagorio S., Børge Johannesen T., Marcos-Gragera R., Francisci S., EUROCARE Working Group Survival of European patients with central nervous system tumors. Int. J. Cancer. 2012;131:173–185. - PubMed
-
- Bao S., Wu Q., McLendon R.E., Hao Y., Shi Q., Hjelmeland A.B., Dewhirst M.W., Bigner D.D., Rich J.N. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature. 2006;444:756–760. - PubMed
-
- Bovenberg M.S., Degeling M.H., Tannous B.A. Advances in stem cell therapy against gliomas. Trends Mol. Med. 2013;19:281–291. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

