Insights into the Functions of LncRNAs in Drosophila
- PMID: 31546813
- PMCID: PMC6770079
- DOI: 10.3390/ijms20184646
Insights into the Functions of LncRNAs in Drosophila
Abstract
Long non-coding RNAs (lncRNAs) are a class of non-coding RNAs longer than 200 nucleotides (nt). LncRNAs have high spatiotemporal specificity, and secondary structures have been preserved throughout evolution. They have been implicated in a range of biological processes and diseases and are emerging as key regulators of gene expression at the epigenetic, transcriptional, and post-transcriptional levels. Comparative analyses of lncRNA functions among multiple organisms have suggested that some of their mechanisms seem to be conserved. Transcriptome studies have found that some Drosophila lncRNAs have highly specific expression patterns in embryos, nerves, and gonads. In vivo studies of lncRNAs have revealed that dysregulated expression of lncRNAs in Drosophila may result in impaired embryo development, impaired neurological and gonadal functions, and poor stress resistance. In this review, we summarize the epigenetic, transcriptional, and post-transcriptional mechanisms of lncRNAs and mainly focus on recent insights into the transcriptome studies and biological functions of lncRNAs in Drosophila.
Keywords: Drosophila; biological function; long non-coding RNAs; mechanism; transcriptome.
Conflict of interest statement
The authors declare no conflicts of interest.
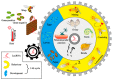
Figures

Similar articles
-
The various and shared roles of lncRNAs during development.Dev Dyn. 2019 Nov;248(11):1059-1069. doi: 10.1002/dvdy.108. Epub 2019 Sep 13. Dev Dyn. 2019. PMID: 31454122 Review.
-
Genome-wide identification and developmental expression profiling of long noncoding RNAs during Drosophila metamorphosis.Sci Rep. 2016 Mar 21;6:23330. doi: 10.1038/srep23330. Sci Rep. 2016. PMID: 26996731 Free PMC article.
-
Long noncoding RNAs in organogenesis: making the difference.Trends Genet. 2015 Jun;31(6):329-35. doi: 10.1016/j.tig.2015.02.002. Epub 2015 Mar 2. Trends Genet. 2015. PMID: 25743487 Review.
-
Long non-coding RNA discovery across the genus anopheles reveals conserved secondary structures within and beyond the Gambiae complex.BMC Genomics. 2015 Apr 23;16(1):337. doi: 10.1186/s12864-015-1507-3. BMC Genomics. 2015. PMID: 25903279 Free PMC article.
-
Comparative Expression Dynamics of Intergenic Long Noncoding RNAs in the Genus Drosophila.Genome Biol Evol. 2016 Jun 27;8(6):1839-58. doi: 10.1093/gbe/evw116. Genome Biol Evol. 2016. PMID: 27189981 Free PMC article.
Cited by
-
Genome-Wide Identification of Long Noncoding RNA and Their Potential Interactors in ISWI Mutants.Int J Mol Sci. 2022 Jun 2;23(11):6247. doi: 10.3390/ijms23116247. Int J Mol Sci. 2022. PMID: 35682924 Free PMC article.
-
Hsrω and Other lncRNAs in Neuronal Functions and Disorders in Drosophila.Life (Basel). 2022 Dec 21;13(1):17. doi: 10.3390/life13010017. Life (Basel). 2022. PMID: 36675966 Free PMC article. Review.
-
Diverse Regulatory Manners and Potential Roles of lncRNAs in the Developmental Process of Asian Honey Bee (Apis cerana) Larval Guts.Int J Mol Sci. 2023 Oct 20;24(20):15399. doi: 10.3390/ijms242015399. Int J Mol Sci. 2023. PMID: 37895079 Free PMC article.
-
Modeling Neoplastic Growth in Renal Cell Carcinoma and Polycystic Kidney Disease.Int J Mol Sci. 2021 Apr 10;22(8):3918. doi: 10.3390/ijms22083918. Int J Mol Sci. 2021. PMID: 33920158 Free PMC article. Review.
-
Post-transcriptional regulation of insect metamorphosis and oogenesis.Cell Mol Life Sci. 2020 May;77(10):1893-1909. doi: 10.1007/s00018-019-03361-5. Epub 2019 Nov 13. Cell Mol Life Sci. 2020. PMID: 31724082 Free PMC article. Review.
References
-
- Derrien T., Johnson R., Bussotti G., Tanzer A., Djebali S., Tilgner H., Guernec G., Martin D., Merkel A., Knowles D.G. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012;22:1775–1789. doi: 10.1101/gr.132159.111. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

