Microbiota fingerprints within the oral cavity of cetaceans as indicators for population biomonitoring
- PMID: 31548611
- PMCID: PMC6757053
- DOI: 10.1038/s41598-019-50139-7
Microbiota fingerprints within the oral cavity of cetaceans as indicators for population biomonitoring
Abstract
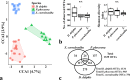
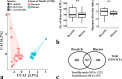
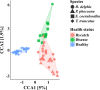
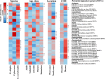
The composition of mammalian microbiota has been related with the host health status. In this study, we assessed the oral microbiome of 3 cetacean species most commonly found stranded in Iberian Atlantic waters (Delphinus delphis, Stenella coeruleoalba and Phocoena phocoena), using 16S rDNA-amplicon metabarcoding. All oral microbiomes were dominated by Proteobacteria, Firmicutes, Bacteroidetes and Fusobacteria bacteria, which were also predominant in the oral cavity of Tursiops truncatus. A Constrained Canonical Analysis (CCA) showed that the major factors shaping the composition of 38 oral microbiomes (p-value < 0.05) were: (i) animal species and (ii) age class, segregating adults and juveniles. The correlation analysis also grouped the microbiomes by animal stranding location and health status. Similar discriminatory patterns were detected using the data from a previous study on Tursiops truncatus, indicating that this correlation approach may facilitate data comparisons between different studies on several cetacean species. This study identified a total of 15 bacterial genera and 27 OTUs discriminating between the observed CCA groups, which can be further explored as microbiota fingerprints to develop (i) specific diagnostic assays for cetacean population conservation and (ii) bio-monitoring approaches to assess the health of marine ecosystems from the Iberian Atlantic basin, using cetaceans as bioindicators.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
The use of DNA barcoding to monitor the marine mammal biodiversity along the French Atlantic coast.Zookeys. 2013 Dec 30;(365):5-24. doi: 10.3897/zookeys.365.5873. eCollection 2013 Dec 30. Zookeys. 2013. PMID: 24453548 Free PMC article.
-
Macroparasites in cetaceans stranded on the northwestern Spanish Atlantic coast.Dis Aquat Organ. 1998 Apr 3;32(3):227-31. doi: 10.3354/dao032227. Dis Aquat Organ. 1998. PMID: 9676247
-
Comparison of the Oral Microbiomes of Canines and Their Owners Using Next-Generation Sequencing.PLoS One. 2015 Jul 2;10(7):e0131468. doi: 10.1371/journal.pone.0131468. eCollection 2015. PLoS One. 2015. PMID: 26134411 Free PMC article.
-
Insights into the human oral microbiome.Arch Microbiol. 2018 May;200(4):525-540. doi: 10.1007/s00203-018-1505-3. Epub 2018 Mar 23. Arch Microbiol. 2018. PMID: 29572583 Review.
-
Oral microbiomes: more and more importance in oral cavity and whole body.Protein Cell. 2018 May;9(5):488-500. doi: 10.1007/s13238-018-0548-1. Epub 2018 May 7. Protein Cell. 2018. PMID: 29736705 Free PMC article. Review.
Cited by
-
Specificity of Stenurus (Metastrongyloidea: Pseudaliidae) infections in odontocetes stranded along the north-west Spanish coast.Int J Parasitol Parasites Wildl. 2022 Sep 13;19:148-154. doi: 10.1016/j.ijppaw.2022.09.002. eCollection 2022 Dec. Int J Parasitol Parasites Wildl. 2022. PMID: 36133958 Free PMC article.
-
Characterization of oral and cloacal microbial communities of wild and rehabilitated loggerhead sea turtles (Caretta caretta).Anim Microbiome. 2021 Sep 3;3(1):59. doi: 10.1186/s42523-021-00120-5. Anim Microbiome. 2021. PMID: 34479653 Free PMC article.
-
Microbiome Composition and Function in Aquatic Vertebrates: Small Organisms Making Big Impacts on Aquatic Animal Health.Front Microbiol. 2021 Mar 11;12:567408. doi: 10.3389/fmicb.2021.567408. eCollection 2021. Front Microbiol. 2021. PMID: 33776947 Free PMC article. Review.
-
Wild whale faecal samples as a proxy of anthropogenic impact.Sci Rep. 2021 Mar 12;11(1):5822. doi: 10.1038/s41598-021-84966-4. Sci Rep. 2021. PMID: 33712645 Free PMC article.
-
Ecology, Not Host Phylogeny, Shapes the Oral Microbiome in Closely Related Species.Mol Biol Evol. 2022 Dec 5;39(12):msac263. doi: 10.1093/molbev/msac263. Mol Biol Evol. 2022. PMID: 36472532 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

