Smad7 Controls Immunoregulatory PDL2/1-PD1 Signaling in Intestinal Inflammation and Autoimmunity
- PMID: 31553906
- PMCID: PMC6925592
- DOI: 10.1016/j.celrep.2019.07.065
Smad7 Controls Immunoregulatory PDL2/1-PD1 Signaling in Intestinal Inflammation and Autoimmunity
Abstract
Smad7, a negative regulator of TGF-β signaling, has been implicated in the pathogenesis and treatment of inflammatory bowel diseases (IBDs), including Crohn's disease (CD) and ulcerative colitis (UC). Here, we found that Smad7 mediates intestinal inflammation by limiting the PDL2/1-PD1 axis in dendritic cells (DCs) and CD4+T cells. Smad7 deficiency in DCs promotes TGF-β responsiveness and the co-inhibitory molecules PDL2/1 on DCs, and it further imprints T cell-PD1 signaling to promote Treg differentiation. DC-specific Smad7 deletion mitigates DSS-induced colitis by inducing CD103+PDL2/1+DCs and Tregs. In addition, Smad7 deficiency in CD4+T cells promotes PD1 and PD1-induced Tregs in vitro. The transfer of Smad7-deficient CD4+T cells enhances Tregs in vivo and protects against T cell-mediated colitis. Furthermore, Smad7 antisense ameliorates DSS-induced UC, increasing TGF-β and PDL2/1-PD1 signaling. Enhancing PD1 signaling directly via Fc-fused PDL2/1 is also beneficial. Our results identify how Smad7 mediates intestinal inflammation and leverages these pathways therapeutically, providing additional strategies for IBD intervention.
Keywords: CD103; DC; IBD; PD1; PDL1; PDL2; Smad7; TGF-β; Treg; UC; dendritic cell; inflammatory bowel disease; ulcerative colitis.
Copyright © 2019 The Author(s). Published by Elsevier Inc. All rights reserved.
Figures



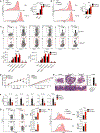

References
-
- Afrakhte M, Morén A, Jossan S, Itoh S, Sampath K, Westermark B, Heldin CH, Heldin NE, and ten Dijke P (1998). Induction of inhibitory Smad6 and Smad7 mRNA by TGF-beta family members. Biochem. Biophys. Res. Commun 249, 505–511. - PubMed
-
- Ajay AK, Kim TM, Ramirez-Gonzalez V, Park PJ, Frank DA, and Vaidya VS (2014). A bioinformatics approach identifies signal transducer and activator of transcription-3 and checkpoint kinase 1 as upstream regulators of kidney injury molecule-1 after kidney injury. J. Am. Soc. Nephrol 25, 105–118. - PMC - PubMed
-
- Baugé C, Attia J, Leclercq S, Pujol JP, Galéra P, and Boumédiene K (2008). Interleukin-1beta up-regulation of Smad7 via NF-kappaB activation in human chondrocytes. Arthritis Rheum. 58, 221–226. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

