Modulation of M2 macrophage polarization by the crosstalk between Stat6 and Trim24
- PMID: 31554795
- PMCID: PMC6761150
- DOI: 10.1038/s41467-019-12384-2
Modulation of M2 macrophage polarization by the crosstalk between Stat6 and Trim24
Abstract
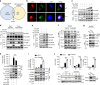
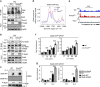
Stat6 is known to drive macrophage M2 polarization. However, how macrophage polarization is fine-tuned by Stat6 is poorly understood. Here, we find that Lys383 of Stat6 is acetylated by the acetyltransferase CREB-binding protein (CBP) during macrophage activation to suppress macrophage M2 polarization. Mechanistically, Trim24, a CBP-associated E3 ligase, promotes Stat6 acetylation by catalyzing CBP ubiquitination at Lys119 to facilitate the recruitment of CBP to Stat6. Loss of Trim24 inhibits Stat6 acetylation and thus promotes M2 polarization in both mouse and human macrophages, potentially compromising antitumor immune responses. By contrast, Stat6 mediates the suppression of TRIM24 expression in M2 macrophages to contribute to the induction of an immunosuppressive tumor niche. Taken together, our findings establish Stat6 acetylation as an essential negative regulatory mechanism that curtails macrophage M2 polarization.
Conflict of interest statement
The authors declare no competing interests.
Figures






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

