Synergy between B cell receptor/antigen uptake and MHCII peptide editing relies on HLA-DO tuning
- PMID: 31554902
- PMCID: PMC6761166
- DOI: 10.1038/s41598-019-50455-y
Synergy between B cell receptor/antigen uptake and MHCII peptide editing relies on HLA-DO tuning
Abstract
B cell receptors and surface-displayed peptide/MHCII complexes constitute two key components of the B-cell machinery to sense signals and communicate with other cell types during antigen-triggered activation. However, critical pathways synergizing antigen-BCR interaction and antigenic peptide-MHCII presentation remain elusive. Here, we report the discovery of factors involved in establishing such synergy. We applied a single-cell measure coupled with super-resolution microscopy to investigate the integrated function of two lysosomal regulators for peptide loading, HLA-DM and HLA-DO. In model cell lines and human tonsillar B cells, we found that tunable DM/DO stoichiometry governs DMfree activity for exchange of placeholder CLIP peptides with high affinity MHCII ligands. Compared to their naïve counterparts, memory B cells with less DMfree concentrate a higher proportion of CLIP/MHCII in lysosomal compartments. Upon activation mediated by high affinity BCR, DO tuning is synchronized with antigen internalization and rapidly potentiates DMfree activity to optimize antigen presentation for T-cell recruitment.
Conflict of interest statement
The authors declare no competing interests.
Figures

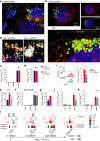




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

