Selective Autophagy of Peroxisomes in Plants: From Housekeeping to Development and Stress Responses
- PMID: 31555306
- PMCID: PMC6722239
- DOI: 10.3389/fpls.2019.01021
Selective Autophagy of Peroxisomes in Plants: From Housekeeping to Development and Stress Responses
Abstract
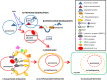
Peroxisomes are dynamic organelles involved in multiple functions, including oxygen and nitrogen reactive species metabolism. In plants, these organelles have a close relationship with chloroplasts and mitochondria, characterized by intense metabolic activity and signal transduction. Peroxisomes undergo rapid changes in size, morphology, and abundance depending on the plant development stage and environmental conditions. As peroxisomes are essential not only for redox homeostasis but also for sensing stress, signaling transduction, and cell survival, their formation and degradation need to be rigorously regulated. In this review, new insights into the regulation of plant peroxisomes are briefly described, with a particular emphasis on pexophagy components and their regulation.
Keywords: autophogy-related genes; peroxins; pexophagy; plants; reactive oxygen species.
Figures
Similar articles
-
Autophagic Machinery of Plant Peroxisomes.Int J Mol Sci. 2019 Sep 25;20(19):4754. doi: 10.3390/ijms20194754. Int J Mol Sci. 2019. PMID: 31557865 Free PMC article. Review.
-
Review: Selective degradation of peroxisome by autophagy in plants: Mechanisms, functions, and perspectives.Plant Sci. 2018 Sep;274:485-491. doi: 10.1016/j.plantsci.2018.06.026. Epub 2018 Jul 2. Plant Sci. 2018. PMID: 30080638 Review.
-
The progression of peroxisomal degradation through autophagy requires peroxisomal division.Autophagy. 2014 Apr;10(4):652-61. doi: 10.4161/auto.27852. Epub 2014 Jan 21. Autophagy. 2014. PMID: 24451165 Free PMC article.
-
Peroxisomes: versatile organelles with diverse roles in plants.New Phytol. 2020 Feb;225(4):1410-1427. doi: 10.1111/nph.16134. Epub 2019 Sep 26. New Phytol. 2020. PMID: 31442305 Review.
-
Pexophagy and peroxisomal protein turnover in plants.Biochim Biophys Acta. 2016 May;1863(5):999-1005. doi: 10.1016/j.bbamcr.2015.09.005. Epub 2015 Sep 5. Biochim Biophys Acta. 2016. PMID: 26348128 Free PMC article. Review.
Cited by
-
Exploring the Contribution of Autophagy to the Excess-Sucrose Response in Arabidopsis thaliana.Int J Mol Sci. 2022 Mar 31;23(7):3891. doi: 10.3390/ijms23073891. Int J Mol Sci. 2022. PMID: 35409249 Free PMC article.
-
Evaluation of protein's interaction and the regulatory network of some drought-responsive genes in Canola under drought and re-watering conditions.Physiol Mol Biol Plants. 2023 Aug;29(8):1085-1102. doi: 10.1007/s12298-023-01345-1. Epub 2023 Sep 1. Physiol Mol Biol Plants. 2023. PMID: 37829706 Free PMC article.
-
Emerging Roles of the Selective Autophagy in Plant Immunity and Stress Tolerance.Int J Mol Sci. 2020 Aug 31;21(17):6321. doi: 10.3390/ijms21176321. Int J Mol Sci. 2020. PMID: 32878263 Free PMC article. Review.
-
Peroxisomal Metabolism and Dynamics at the Crossroads Between Stimulus Perception and Fast Cell Responses to the Environment.Front Cell Dev Biol. 2020 Jun 26;8:505. doi: 10.3389/fcell.2020.00505. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 32676503 Free PMC article. No abstract available.
-
Peroxisomes as redox-signaling nodes in intracellular communication and stress responses.Plant Physiol. 2021 May 27;186(1):22-35. doi: 10.1093/plphys/kiab060. Plant Physiol. 2021. PMID: 33587125 Free PMC article. Review.
References
-
- Alvarez C., Garcia I., Moreno I., Perez-Perez M. E., Crespo J. L., Romero L. C., et al. (2012). Cysteine-generated sulfide in the cytosol negatively regulates autophagy and modulates the transcriptional profile in Arabidopsis . Plant Cell 24 (11), 4621–4634. 10.1105/tpc.112.105403 - DOI - PMC - PubMed
-
- Barros J. A. S., Cavalcanti J. H. F., Medeiros D. B., Nunes-Nesi A., Avin-Wittenberg T., Fernie A. R., et al. (2017). Commonalities and differences in plants deficient in autophagy and alternative pathways of respiration on response to extended darkness. Plant Signal. Behav. 12 (11), e1377877. 10.1080/15592324.2017.1377877 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Research Materials

