Comprehensive Landscape of Active Deubiquitinating Enzymes Profiled by Advanced Chemoproteomics
- PMID: 31555637
- PMCID: PMC6727631
- DOI: 10.3389/fchem.2019.00592
Comprehensive Landscape of Active Deubiquitinating Enzymes Profiled by Advanced Chemoproteomics
Abstract
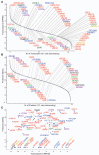
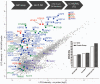
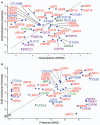
Enzymes that bind and process ubiquitin, a small 76-amino-acid protein, have been recognized as pharmacological targets in oncology, immunological disorders, and neurodegeneration. Mass spectrometry technology has now reached the capacity to cover the proteome with enough depth to interrogate entire biochemical pathways including those that contain DUBs and E3 ligase substrates. We have recently characterized the breast cancer cell (MCF7) deep proteome by detecting and quantifying ~10,000 proteins, and within this data set, we can detect endogenous expression of 65 deubiquitylating enzymes (DUBs), whereas matching transcriptomics detected 78 DUB mRNAs. Since enzyme activity provides another meaningful layer of information in addition to the expression levels, we have combined advanced mass spectrometry technology, pre-fractionation, and more potent/selective ubiquitin active-site probes with propargylic-based electrophiles to profile 74 DUBs including distinguishable isoforms for 5 DUBs in MCF7 crude extract material. Competition experiments with cysteine alkylating agents and pan-DUB inhibitors combined with probe labeling revealed the proportion of active cellular DUBs directly engaged with probes by label-free quantitative (LFQ) mass spectrometry. This demonstrated that USP13, 39, and 40 are non-reactive to probe, indicating restricted enzymatic activity under these cellular conditions. Our extended chemoproteomics workflow increases depth of covering the active DUBome, including isoform-specific resolution, and provides the framework for more comprehensive cell-based small-molecule DUB selectivity profiling.
Keywords: chemical biology; deubiquitylating enzymes; isoforms; mass spectrometry; proteomics; ubiquitin specific proteases.
Figures


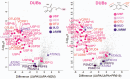
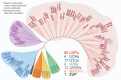
References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

