Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome
- PMID: 31561566
- PMCID: PMC6801519
- DOI: 10.3390/ijms20194788
Complete Sequence, Multichromosomal Architecture and Transcriptome Analysis of the Solanum tuberosum Mitochondrial Genome
Abstract
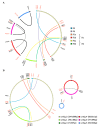
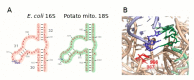
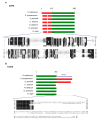
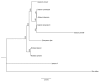
Mitochondrial genomes (mitogenomes) in higher plants can induce cytoplasmic male sterility and be somehow involved in nuclear-cytoplasmic interactions affecting plant growth and agronomic performance. They are larger and more complex than in other eukaryotes, due to their recombinogenic nature. For most plants, the mitochondrial DNA (mtDNA) can be represented as a single circular chromosome, the so-called master molecule, which includes repeated sequences that recombine frequently, generating sub-genomic molecules in various proportions. Based on the relevance of the potato crop worldwide, herewith we report the complete mtDNA sequence of two S. tuberosum cultivars, namely Cicero and Désirée, and a comprehensive study of its expression, based on high-coverage RNA sequencing data. We found that the potato mitogenome has a multi-partite architecture, divided in at least three independent molecules that according to our data should behave as autonomous chromosomes. Inter-cultivar variability was null, while comparative analyses with other species of the Solanaceae family allowed the investigation of the evolutionary history of their mitogenomes. The RNA-seq data revealed peculiarities in transcriptional and post-transcriptional processing of mRNAs. These included co-transcription of genes with open reading frames that are probably expressed, methylation of an rRNA at a position that should impact translation efficiency and extensive RNA editing, with a high proportion of partial editing implying frequent mis-targeting by the editing machinery.
Keywords: RNA editing; Solanaceae family; comparative genomics; mitochondria; mtDNA; multichromosomal structure; potato; repeated sequences.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
The complete chloroplast genome sequences of Solanum tuberosum and comparative analysis with Solanaceae species identified the presence of a 241-bp deletion in cultivated potato chloroplast DNA sequence.Plant Cell Rep. 2006 Dec;25(12):1369-79. doi: 10.1007/s00299-006-0196-4. Epub 2006 Jul 12. Plant Cell Rep. 2006. PMID: 16835751
-
Mitochondrial gene editing and allotopic expression unveil the role of orf125 in the induction of male fertility in some Solanum spp. hybrids and in the evolution of the common potato.Plant Biotechnol J. 2025 May;23(5):1862-1875. doi: 10.1111/pbi.70012. Epub 2025 Mar 22. Plant Biotechnol J. 2025. PMID: 40119623 Free PMC article.
-
Breaking the limits - multichromosomal structure of an early eudicot Pulsatilla patens mitogenome reveals extensive RNA-editing, longest repeats and chloroplast derived regions among sequenced land plant mitogenomes.BMC Plant Biol. 2022 Mar 9;22(1):109. doi: 10.1186/s12870-022-03492-1. BMC Plant Biol. 2022. PMID: 35264098 Free PMC article.
-
Multichromosomal structure of the onion mitochondrial genome and a transcript analysis.Mitochondrion. 2019 May;46:179-186. doi: 10.1016/j.mito.2018.05.001. Epub 2018 Jul 10. Mitochondrion. 2019. PMID: 30006008
-
The complete mitogenome assemblies of 10 diploid potato clones reveal recombination and overlapping variants.DNA Res. 2021 Aug 25;28(4):dsab009. doi: 10.1093/dnares/dsab009. DNA Res. 2021. PMID: 34254134 Free PMC article.
Cited by
-
Comparative analysis of the complete mitogenomes of Camellia sinensis var. sinensis and C. sinensis var. assamica provide insights into evolution and phylogeny relationship.Front Plant Sci. 2024 Aug 22;15:1396389. doi: 10.3389/fpls.2024.1396389. eCollection 2024. Front Plant Sci. 2024. PMID: 39239196 Free PMC article.
-
Characterizing complete mitochondrial genome of Aquilegia amurensis and its evolutionary implications.BMC Plant Biol. 2024 Feb 28;24(1):142. doi: 10.1186/s12870-024-04844-9. BMC Plant Biol. 2024. PMID: 38413922 Free PMC article.
-
Comparative Analysis of Tylosema esculentum Mitochondrial DNA Revealed Two Distinct Genome Structures.Biology (Basel). 2023 Sep 16;12(9):1244. doi: 10.3390/biology12091244. Biology (Basel). 2023. PMID: 37759643 Free PMC article.
-
Assembly of the Complete Mitochondrial Genome of Chinese Plum (Prunus salicina): Characterization of Genome Recombination and RNA Editing Sites.Genes (Basel). 2021 Dec 10;12(12):1970. doi: 10.3390/genes12121970. Genes (Basel). 2021. PMID: 34946920 Free PMC article.
-
Sequence of the Mitochondrial Genome of Lactuca virosa Suggests an Unexpected Role in Lactuca sativa's Evolution.Front Plant Sci. 2021 Jul 26;12:697136. doi: 10.3389/fpls.2021.697136. eCollection 2021. Front Plant Sci. 2021. PMID: 34381482 Free PMC article.
References
-
- Sloan D.B., Alverson A.J., Chuckalovcak J.P., Wu M., McCauley D.E., Palmer J.D., Taylor D.R. Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol. 2012;10:e1001241. doi: 10.1371/journal.pbio.1001241. - DOI - PMC - PubMed
-
- Rodríguez-Moreno L., González V.M., Benjak A., Martí M.C., Puigdomènech P., Aranda M.A., Garcia-Mas J. Determination of the melon chloroplast and mitochondrial genome sequences reveals that the largest reported mitochondrial genome in plants contains a significant amount of DNA having a nuclear origin. BMC Genom. 2011;12:424. doi: 10.1186/1471-2164-12-424. - DOI - PMC - PubMed
-
- Morley S.A., Nielsen B.L. Plant mitochondrial DNA. Molecules. 2017;15:17. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

