The genomic landscape of metastatic breast cancer highlights changes in mutation and signature frequencies
- PMID: 31570896
- PMCID: PMC6858873
- DOI: 10.1038/s41588-019-0507-7
The genomic landscape of metastatic breast cancer highlights changes in mutation and signature frequencies
Abstract
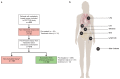
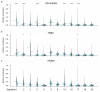
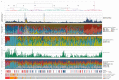
The whole-genome sequencing of prospectively collected tissue biopsies from 442 patients with metastatic breast cancer reveals that, compared to primary breast cancer, tumor mutational burden doubles, the relative contributions of mutational signatures shift and the mutation frequency of six known driver genes increases in metastatic breast cancer. Significant associations with pretreatment are also observed. The contribution of mutational signature 17 is significantly enriched in patients pretreated with fluorouracil, taxanes, platinum and/or eribulin, whereas the de novo mutational signature I identified in this study is significantly associated with pretreatment containing platinum-based chemotherapy. Clinically relevant subgroups of tumors are identified, exhibiting either homologous recombination deficiency (13%), high tumor mutational burden (11%) or specific alterations (24%) linked to sensitivity to FDA-approved drugs. This study provides insights into the biology of metastatic breast cancer and identifies clinically useful genomic features for the future improvement of patient management.
Conflict of interest statement
Figures


References
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

